Amyloid β protein
Amyloid beta (Aβ or Abeta) denotes peptides of 36–43 amino acids that are the main component of the amyloid plaques found in the brains of people with Alzheimer's disease. The peptides derive from the amyloid precursor protein (APP), which is cleaved by beta secretase and gamma secretase to yield Aβ. Aβ molecules can aggregate to form flexible soluble oligomers which may exist in several forms. It is now believed that certain misfolded oligomers (known as "seeds") can induce other Aβ molecules to also take the misfolded oligomeric form, leading to a chain reaction akin to a prion infection. The oligomers are toxic to nerve cells. The other protein implicated in Alzheimer's disease, tau protein, also forms such prion-like misfolded oligomers, and there is some evidence that misfolded Aβ can induce tau to misfold.
The normal function of Aβ is not well understood. Though some animal studies have shown that the absence of Aβ does not lead to any obvious loss of physiological function, several potential activities have been discovered for Aβ, including activation of kinase enzymes, protection against oxidative stress, regulation of cholesterol transport, functioning as a transcription factor, and anti-microbial activity (potentially associated with Aβ's pro-inflammatory activity).
The glymphatic system clears metabolic waste from the mammalian brain, and in particular amyloid beta. Indeed, a number of proteases have been implicated by both genetic and biochemical studies as being responsible for the recognition and degradation of amyloid beta; these include insulin degrading enzyme.and presequence protease. The rate of removal is significantly increased during sleep. However, the significance of the lymphatic system in Aβ clearance in Alzheimer's disease is unknown.
Targets for Amyloid β protein
Products for Amyloid β protein
- Cat.No. Product Name Information
-
GC37984
β-Amyloid (1-42), rat
β-Amyloid (1-42), rat is a 42-aa peptide, shows cytotoxic effect on acute hippocampal slices, and used in the research of Alzheimer's disease.

-
GC61988
β-amyloid (12-28) (TFA)

-
GC37986
β-Amyloid 1-17
β-Amyloid 1-17 is a peptide of β-Amyloid, stabilizes the fibres and plays a role in Aβ fibre formation.

-
GC37987
β-Amyloid 1-20
β-Amyloid 1-20 consists of amino acids 1 to 20 of beta amyloid protein.

-
GC37990
β-Amyloid 1-34
β-Amyloid 1-34 is a β-Amyloid peptide consists of 34 amino acid.

-
GC37993
β-Amyloid 1-9
β-Amyloid 1-9, an N-terminal fragment of beta amyloid, consists of amino acid residues 1 to 9.
-
GC37985
β-Amyloid 11-22
β-Amyloid 11-22 is a peptide fragment of β-Amyloid.
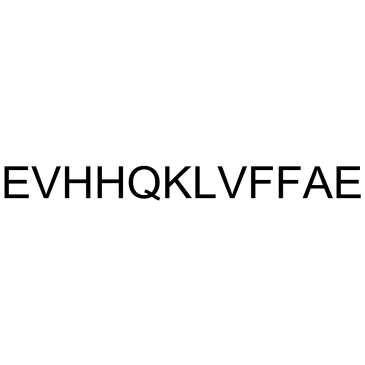
-
GC37988
β-Amyloid 12-20
β-Amyloid 12-20 is a peptide fragment of β-Amyloid.
-
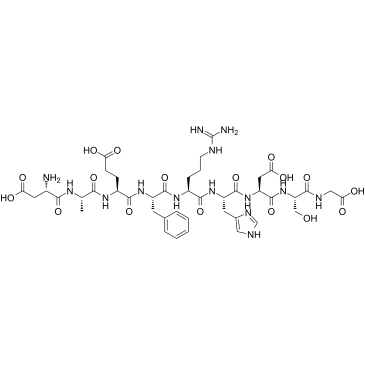
GC37989
β-Amyloid 13-27
β-Amyloid 13-27 is a peptide consisting of amino acid of 13 to 27 of beta amyloid protein.
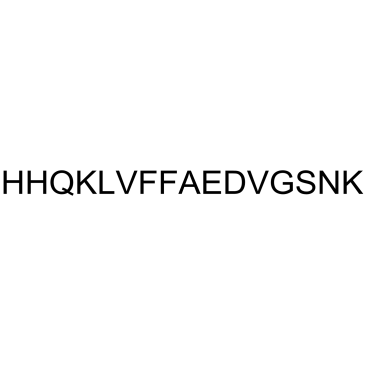
-
GC39466
β-Amyloid 15-21

-
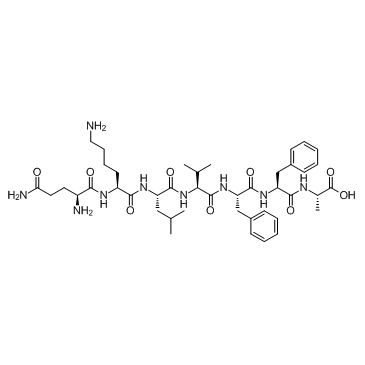
GC37991
β-Amyloid 15-21
-
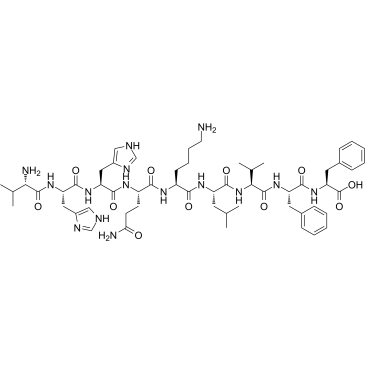
GC37992
β-Amyloid 18-28
β-Amyloid 18-28 is a peptide fragment of β-Amyloid.
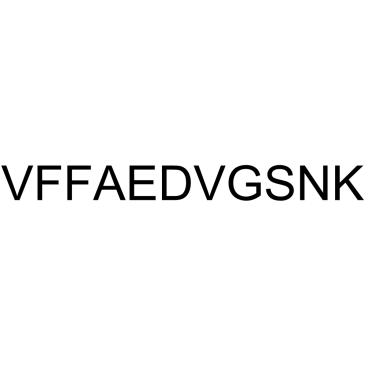
-
GC37994
β-Amyloid 22-40
β-Amyloid 22-40 is a peptide fragment of β-Amyloid.
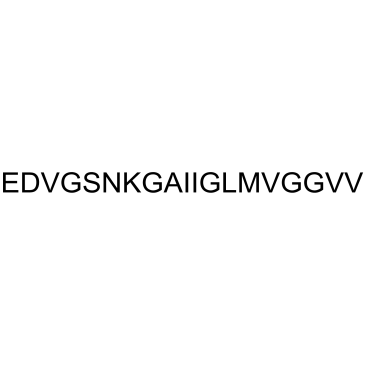
-
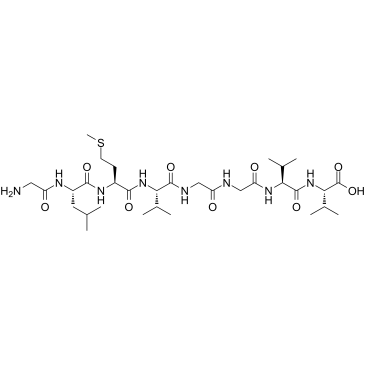
GC37995
β-Amyloid 33-40
β-Amyloid 33-40 is a peptide consisting of amino acid of 33 to 40 of beta amyloid protein.
-
GC37996
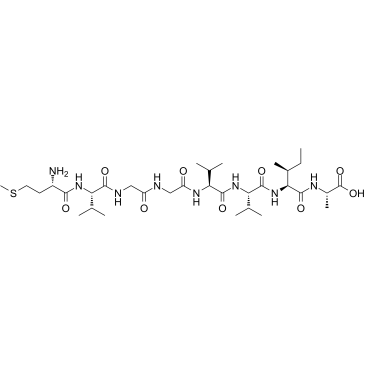
β-Amyloid 35-42
β-Amyloid 35-42 is a peptide consisting of amino acid of 35 to 42 of beta amyloid protein.
-
GC37997
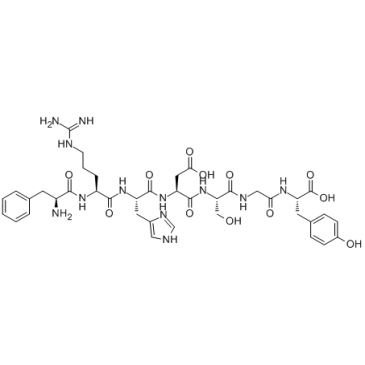
β-Amyloid 4-10
β-Amyloid 4-10 is an epitope for the polyclonal anti-Aβ(1-42) antibody, reduces amyloid deposition in a transgenic Alzheimer disease mouse model.
-
GC37998
β-Amyloid Protein Precursor 770 135-155
β-Amyloid Protein Precursor 770 135-155 is a peptide of amyloid precursor protein isoform (APP 770).

-
GC34242
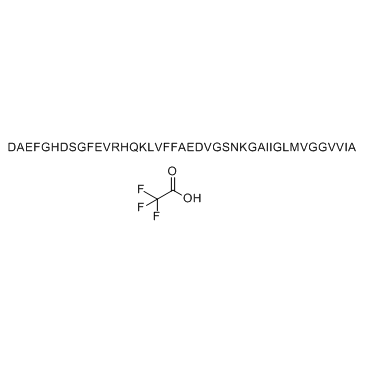
β-Amyloid (1-42), rat TFA
-
GC31146
β-Amyloid (10-35), amide
β-Amyloid (10-35), amide is composed of 26 aa (10-35 residues of the Aβ peptide) and is the primary component of the amyloid plaques of Alzheimer's disease.

-
GC31129
β-Amyloid 1-16 (Amyloid β-Protein (1-16))
β-Amyloid 1-16 (Amyloid β-Protein (1-16)) is a β-Amyloid protein fragment involved in metal binding.

-
GC31171
β-Amyloid 1-28 (Amyloid β-Protein (1-28))
β-Amyloid 1-28 (Amyloid β-Protein (1-28)) is a β-Amyloid protein fragment involved in metal binding.
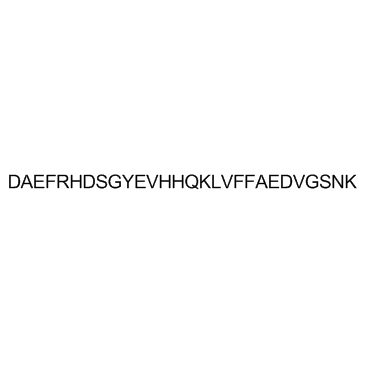
-
GC34391
β-Amyloid 15-21 (Beta-Amyloid (15-21))
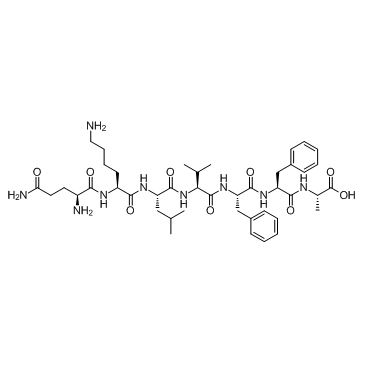
-
GC30325
β-Amyloid 22-35 (Amyloid β-Protein (22-35))
β-Amyloid 22-35 (Amyloid β-Protein 22-35), the residues 22-35 fragment ofβ-amyloid protein, has a cytotoxic effect on cultured neurons from the rat hippocampus in serum-free medium.

-
GC31137
β-Amyloid 29-40 (Amyloid beta-protein(29-40))
β-Amyloid 29-40 (Amyloid beta-protein(29-40)) is a fragment of Amyloid-β peptide.

-
GC31179
β-Amyloid 31-35
β-Amyloid 31-35 is the shortest sequence of native Amyloid-β peptide that retains neurotoxic activity.
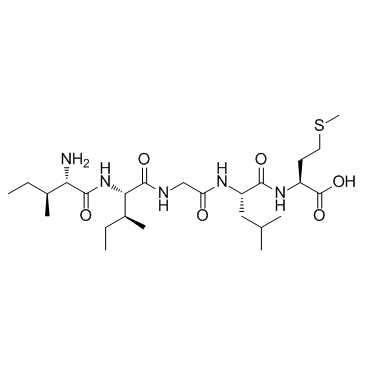
-
GA20024
(7-Diethylaminocoumarin-3-yl)carbonyl-Amyloid β-Protein (1-40)
Amyloid β-protein (1-40) that is N-terminally modified with the fluorescent dye (7-diethylaminocoumarin-3-yl)carbonyl (DAC or DEAC). This derivative can be utilized to assess the binding properties of amyloid β-protein (1-40) for various membranes since it behaves very similar to the native peptide. In aqueous environments the fluorophore is almost non-fluorescent whereas binding to membranes results in an increase in fluorescence intensity (Λex = 430 nm, Λem = 470 nm). Increases in the GM1 ganglioside and cholesterol content in the lipid bilayers facilitated the binding of this peptide. For phosphatidylcholine and phosphatidylserine no affinity was observed.

-
GA20029
(Arg¹³)-Amyloid β-Protein (1-40)
H13R, a mutation in the metal-binding region of Abeta reduces its copper-mediated toxicity. The native rodent sequence containing an arginine at this position is more tolerant to metals than the human amyloid peptide.

-
GA20030
(Arg⁶)-Amyloid β-Protein (1-40)
The English (H6R) mutation of β-amyloid peptides accelerates fibrillation without increasing protofibril formation. Ono et al. showed that the English and Tottori mutations alter Abeta assembly at its earliest stages, monomer folding and oligomerization, and produce oligomers that are more toxic to cultured neuronal cells than are wild type oligomers.
The exchange of His? by Arg influences the structure of the Cu(II) complex formed by Aβ peptides.

-
GA20038
(Asn²³)-Amyloid β-Protein (1-40)
The Iowa (D23N) mutant of Aβ 40 considerably more rapidly assembles in solution to form fibrils than the WT Aβ sequence. These fibrils also show a different structure, which could be responsible for their increased toxicity.

-
GA20039
(Asn⁶⁷⁰,Leu⁶⁷¹)-Amyloid β/A4 Protein Precursor₇₇₀ (667-675)
SEVNLDAEF corresponds to the mutant junctional sequence of the amyloid precursor protein (APP) found in a Swedish family with early-onset Alzheimer's disease, therefore referred to as the 'Swedish' mutation (K670N/M671L). The peptide has been used for assaying cleavage at leucine-aspartate by cathepsin G and chymotrypsin, whereas neither cathepsin B, D nor L generated any products.

-
GA20040
(Asn⁶⁷⁰,Leu⁶⁷¹)-Amyloid β/A4 Protein Precursor₇₇₀ (667-676)
This peptide substrate corresponds to the 'Swedish' Lys-Met/Asn-Leu (K670N/M671L) mutation of the amyloid precursor protein (APP) β-secretase cleavage site. It has been used for assaying β-secretase activity.

-
GA20041
(Asn⁶⁷⁰,Sta⁶⁷¹,Val⁶⁷²)-Amyloid β/A4 Protein Precursor₇₇₀ (662-675)
Amyloid precursor protein (APP) β-secretase from human brain cleaves full-length APP at the amino terminus of the amyloid β-protein (Aβ) sequence, thus leading to the generation and extracellular release of β-cleaved soluble APP and a corresponding cell-associated carboxy-terminal fragment. The subsequent cleavage of the C-terminal fragment by γ-secretase(s) leads to the formation of Aβ. This new peptide represents a potent substrate analog inhibitor of APP β-secretase with IC?? = 30 nM.

-
GA20042
(Asn⁷)-Amyloid β-Protein (1-40)
The Tottori (D7N) mutation of β-amyloid peptides accelerates fibrillation without increasing protofibril formation. Ono et al. showed that the English and Tottori mutations alter Abeta assembly at its earliest stages, monomer folding and oligomerization, and produce oligomers that are more toxic to cultured neuronal cells than are wild type oligomers.

-
GA20045
(Asp³⁷)-Amyloid β-Protein (1-42)
The G37D mutant does not show the aggregation behavior of WT Abeta42 nor its neurotoxicity.

-
GA20053
(Cys²⁶)-Amyloid β-Protein (1-40)
Aβ40 S26C has been used for generating the covalently linked Aβ40 homodimer. Dimerization can be easily reverted by reducing the soluble dimer with thiols as β-mercaptoethanol. Aβ40 S26C is perfectly suited for labeling with fluorescent tags

-
GA20052
(Cys²⁶)-Amyloid β-Protein (1-40) (Dimer)
Dimer of H-7402.

-
GA20050
(Cys⁰)-Amyloid β-Protein (1-40)
Cys-Aβ1-40 can be easily and selectively modified, labeled, coupled to carriers e.g. by maleimide chemistry without affecting the sequences involved in fibril formation. The free mercapto moiety of the peptide adheres to gold surfaces.

-
GA20095
(Des-Glu²²)-Amyloid β-Protein (1-40)
The Osaka mutation was the first deletion-type mutation to be identified in APP and Aβ.
The Aβ E22delta mutant is more resistant to degradation by two major Aβ-degrading enzymes, neprilysin and insulin-degrading enzyme. Synthetic mutant Aβ showed unusual aggregation properties with enhanced oligomerization but no fibrillization. It also inhibited hippocampal long-term potentiation more efficiently than wild-type Aβ. A transgenic mouse model containing APP with the E693delta mutation has been developed. APP(OSK)-Tg mice exhibit intraneuronal Aβ E22delta oligomers and memory impairment as early as eight months of age.

-
GA20096
(Des-Glu²²)-Amyloid β-Protein (1-42)
The Osaka (E22delta) mutation of Amyloid β promotes β-sheet transformation, radical production, and synaptotoxicity, but not neurotoxicity.

-
GA20182
(Gln²²)-Amyloid β-Protein (1-40)
The Dutch mutation (E22Q) of amyloid β-peptide aggregates more readily than the wild-type peptide and the resulting fibrils show increased neurotoxicity. The mutant peptide E22Q induced apoptosis of cerebral endothelial cells at a concentration of 25 μm, whereas WT Aβ 1-40 and the Italian mutant E22K (H-6698) showed no effect.

-
GA20183
(Gln²²)-Amyloid β-Protein (1-42)
The Dutch mutation (E22Q) aggregates more readily than the wild-type sequence. The resulting fibrils show increased neurotoxicity.

-
GA20184
(Gln²²,Asn²³)-Amyloid β-Protein (1-40)
Transgenic mice expressing the vasculotropic Dutch/Iowa (E693Q/D694N) mutant human Aβ precursor protein in brain (Tg-SwDI) accumulate abundant cerebral microvascular fibrillar amyloid deposits and exhibit robust neuroinflammation. In vitro, the doubly mutated Aβ peptides showed an increased propensity to fibrillation and pathogenicity compared to the Dutch and Iowa single mutants.

-
GA20181
(Gln¹¹)-Amyloid β-Protein (1-28)

-
GA20186
(Gln⁹)-Amyloid β-Protein (1-40)

-
GA20199
(Gly²²)-Amyloid β-Protein (1-40)
The highly neurotoxic arctic mutant (E22G) of Aβ has been used to study the mechanisms underlying the formation of soluble and insoluble β-amyloid aggregates. As the wild-type Aβ, the arctic mutant preferably assembles in the presence of GM1 ganglioside.

-
GA20200
(Gly²²)-Amyloid β-Protein (1-42)
The arctic mutant of amyloid β peptide 1-42, in which Glu²² is substituted by Gly, is distinctly more amyloidogenic than the wild-type Aβ 1-42.

-
GA20197
(Gly²¹)-Amyloid β-Protein (1-40)
Contrary to β-amyloid peptides mutated at position 22 (Dutch, Italian, Arctic mutants) the Flemish mutation (A21G) shows a decreased tendency to aggregate and a reduced neurotoxicity. In the studies of Betts and Tsubuki, A21G was degraded significantly more slowly by neprilysin than the wild-type Aβ 1-40 and the E22 mutants. The relative resistance to proteolytic degradation may account for the pathogenicity of the Aβ mutant.

-
GA20198
(Gly²¹)-Amyloid β-Protein (1-42)
The Flemish mutation (A21G) shows a decreased tendency to aggregate and a reduced neurotoxicity. A21G is pathogenic as it is degraded significantly more slowly by neprilysin than WT Abeta42.

-
GA20201
(Gly²⁸,Cys³⁰)-Amyloid β-Protein (1-30) amide

-
GA20244
(Lys²²)-Amyloid β-Protein (1-40)
The Italian mutation of β-amyloid 1-40 (E22K) aggregates more rapidly than the wild-type sequence 1-40. It showed increased neurotoxicity, which (according to a solid-phase NMR-study of Masuda et al.) may be due to the salt bridge formed between Lys²² and Asp²³ in the minor conformer. As the Arctic, Flemish, and Dutch mutants, the Italian mutant is degraded considerably more slowly than wild-type Aβ by neprilysin.

-
GA20245
(Lys²²)-Amyloid β-Protein (1-42)
The Italian mutation (E22K) aggregates more rapidly than the wild-type sequence.

-
GA20242
(Lys¹⁵)-Amyloid β-Protein (15-21)
KKLVFFA contains the KLVFF sequence, which is the minimum sequence binding the full-length amyloid β-protein. It showed improved water solubility compared with KLVFF (H-3682). It can be used as a labeled probe for screening defined sequences in the full-length amyloid β-protein.

-
GA20251
(Met(O)³⁵)-Amyloid β-Protein (1-40)
Oxidation of Met35 attenuates the formation of Aβ40 oligomers.

-
GA20252
(Met(O)³⁵)-Amyloid β-Protein (1-42)
(Met(O)³?)-Amyloid β-protein (1-42) (H-5888), in contrast to Aβ 1-42 (H-1368), has been shown to be non-toxic to 9-11 day-old rat embryonic hippocampal neuronal cultures and not to produce any protein oxidation. It has also been demonstrated that fibril formation is not affected by Met(O)³?. For the Nle analog see H-7308.

-
GA20253
(Met(O)³⁵)-Amyloid β-Protein (25-35)
Sulfoxide of Aβ 25-35.

-
GA20254
(Met(O₂)³⁵)-Amyloid β-Protein (1-42)
Maiti et al. could show that, in contrast to the sulfoxide of Aβ (1-42), the sulfone was as toxic and aggregated as fast as wild-type Aβ (1-42).

-
GA20259
(Nle³⁵)-Amyloid β-Protein (1-40)
The reactive thioether of Met³? is crucial for the activity of Aβ 1-40 and Aβ 1-42. Due to the replacement of Met by inert Nle, M35Nle Aβ 1-40 was no longer toxic to cultured hippocampal neurons and had little effect on the level of protein carbonyl residues. The Nle peptide showed the same propensity to aggregate, whereas sulfoxide formation hindered the required conformational transition from random coil to β-sheet.

-
GA20260
(Nle³⁵)-Amyloid β-Protein (1-42)
The thioether of Met³? plays a critical role in the oxidative stress induced by Aβ 1-42 and its neurotoxicity. The norleucine analog Aβ 1-42 M35Nle forms fibrils morphologically indistinguishable from the ones of the native sequence though lacking their neurotoxicity.

-
GA20283
(Pyr³)-Amyloid β-Protein (3-40)
The pyroglutamate-modified amyloid-β peptides derived from Aβ40 (H-7422) and Aβ42 (H-4796) have gained considerable attention as potential key participants in the pathology of Alzheimer's disease (AD) due to their abundance in AD brain, high aggregation propensity, stability, and cellular toxicity. Aβ40 and 42 can be N-terminally truncated by action of cathepsin B. The cyclization of Glu³ is catalyzed by glutaminyl cyclase. Hence, inhibition of these enzymes could be a therapeutic approach to AD.

-
GA20285
(Pyr³)-Amyloid β-Protein (3-42)

-
GA20284
(Pyr³)-Amyloid β-Protein (3-42)
(Pyr³)-Amyloid β-Protein (3-42) was found to be the predominant amyloid β-peptide structure deposited in human brain of Alzheimer's disease and Down's syndrome patients. Therefore, (Pyr³)-Aβ (3-42) is suggested to accumulate in the brain and to trigger the formation of insoluble amyloid β-peptide deposits. Nussbaum et al. studies the Prion-like behaviour and tau-dependent cytotoxicity of the truncated Aβ sequence.

-
GA20282
(Pyr¹¹)-Amyloid β-Protein (11-40)
pEVHHQKLVFFAEDVGSNKGAIIGLMVGGVV, the N-terminally truncated isoform of the amyloid β-protein (Aβ) beginning with a pyroglutamate (Pyr) residue at position 11 was used in experiments studying the generality of fibrillogenesis-related helix formation. Comparing the fibrillogenesis kinetics of many of the most important clinically relevant amyloid β-protein alloforms it could be observed that among these peptides (Pyr¹¹)-amyloid β-protein (11-40) exhibited the greatest retardation of fibrillization rate.

-
GA20309
(Thr²)-Amyloid β-Protein (1-42)
A mutation very close to the β-secretase cleavage site of APP. The Icelandic mutation A2T of Aβ42 turned out to be less pathogenic than the native sequence. The precursor APP A673T was the first APP variant discovered in humans reducing the risk of Alzheimer's disease. A2T as well affects γ-secretase cleavage, the mutant was an inefficient substrate in a cell-based assay of the enzyme.

-
GA20339
(Val²)-Amyloid β-Protein (1-42)
A mutation very close to the β-secretase cleavage site of APP (A673V). Contrary to the protective Icelandic mutation A2T, the recessive A2V mutation may increase the risk of Alzheimer's disease. Cantu et al. observed that APP A673V is associated with the early onset of AD-type dementia in homozygous individuals, whereas it has a protective effect in the heterozygous state.

-
GA20340
(Val²)-Amyloid β-Protein (1-6)

-
GA20341
(Val³⁴)-Amyloid β-Protein (1-40)
Cerebral amyloid angiopathy (CAA) is a common finding in Alzheimer's disease in which amyloid-Aβ vascular deposits are featured in >80% of the cases. Mutations in the positions 21-23 (e.g. Dutch mutation E22Q) are primarily associated with CAA, although they manifest with strikingly different clinical phenotypes: cerebral hemorrhage or dementia. The Piedmont L34V Aβ mutant, located outside this hot spot, shows a similar hemorrhagic phenotype, albeit less aggressive than the widely studied Dutch variant.

-
GA20433
5-FAM-Amyloid β-Protein (1-40)
5-FAM-Amyloid β-Protein (1-40) is a FAM fluorescently-labelledβ-Amyloid (1-40) peptide (Λex=492nm and Λem=518nm).

-
GA20435
5-FAM-Amyloid β-Protein (1-42)

-
GA20434
5-FAM-Amyloid β-Protein (1-42) (scrambled)
Fluorescent dye-labeled inactive control for H-1368, H-6466, H-8146.

-
GC42505
5-FAM-Amyloid-β (1-28) Peptide (human) (trifluoroacetate salt)
5-FAM-Amyloid-β (1-28) peptide is a fluorescently labeled peptide.

-
GC40130
5-FAM-Amyloid-β (1-42) Peptide (human) (trifluoroacetate salt)
5-FAM-Amyloid-β (1-42) peptide is a fluorescently labeled amyloid-β peptide.

-
GA20440
5-TAMRA-Amyloid β-Protein (1-40)
Anderson and Webb could verify using transmission electron microscopy that N-terminal labeling of Aβ40 with TAMRA and other fluorescent dyes does not prevent the formation of protofibrils and amyloid fibrils of various widths.

-
GA20441
5-TAMRA-Amyloid β-Protein (1-42)

-
GA20448
Abz-(Asn⁶⁷⁰,Leu⁶⁷¹)-Amyloid β/A4 Protein Precursor₇₇₀ (669-674)-EDDnp
Intramolecularly quenched fluorescent substrate containing the ortho-aminobenzoyl (Abz) / N-(2,4-dinitrophenyl)ethylenediamine (EDDnp) groups as the donor / acceptor pair. It corresponds to the Swedish-mutated (JMV2236) β-amyloid precursor protein (βAPP) sequence targeted by β-secretase BACE (β-site APP-cleaving activity). This FRET substrate is more selectively cleaved by BACE1 and BACE2 than by cathepsin D, a disintegrin and metalloprotease 10 (ADAM10), tumor necrosis α-converting enzyme (TACE), presenilin-1 (PS1), or presenilin-2 (PS2).

-
GA20452
Abz-Amyloid β/A4 Protein Precursor₇₇₀ (669-674)-EDDnp
Intramolecularly quenched fluorescent substrate containing the ortho-aminobenzoyl (Abz) / N-(2,4-dinitrophenyl)ethylenediamine (EDDnp) groups as the donor / acceptor pair. It mimicks the wild-type (JMV2235) β-amyloid precursor protein (βAPP) sequence targeted by β-secretase BACE (β-site APP-cleaving activity). This FRET substrate is cleaved by BACE1, BACE2, and cathepsin D.

-
GA20453
Abz-Amyloid β/A4 Protein Precursor₇₇₀ (708-715)-Lys(Dnp)-D-Arg-D-Arg-D-Arg amide
A sensitive fluorogenic (FRET) substrate developed for the analysis of γ-secretase from post mortem non-Alzheimer's and Alzheimer's disease human brain isolates.

-
GA20525
Acetyl-(N-Me-Leu¹⁷,N-Me-Phe¹⁹)-Amyloid β-Protein (16-20) amide
Membrane-permeable inhibitor of Aβ (1-40) fibrillogenesis.

-
GA20526
Acetyl-(Pro¹⁸,Asp²¹)-Amyloid β-Protein (17-21) amide
The pentapeptide Ac-LPFFD-NH? (iAβ5p) is an analog of product H-4876 with small chemical modifications which enhance its stability against proteolytic degradation. iAβ5p acts as a β-sheet breaker peptide and crosses the blood-brain barrier at a higher rate than most proteins and peptides known to be selectively taken up by the brain so that it is assumed that the peptide is being specifically transported to the brain. A significant increase in neuronal survival and decrease in brain inflammation associated with the reduction of amyloid plaques in two different transgenic Alzheimer`s disease (AD) models is additionally reported for iAβ5p.

-
GA20535
Acetyl-Amyloid β-Protein (1-6) amide
Experiments using sub-peptides of Aβ42 revealed that the epitope identified by the antibody A8, as described by Ying and coworkers, lies within the 1-6 region of Aβ. The antibody displays high affinity for soluble Aβ42 oligomers in the molecular weight range of 16.5-25 kDa, and detected target antigen in brain sections from senescence-accelerated SAMP 8 mice. Amidated or acetylated and amidated forms of the sequence were used for example for quantitative structure retention relationships (QSRR) experiments. The latter could allow prediction of reversed-phase high-performance liquid chromatography (HPLC) retention of peptides, as reported by Kaliszan and coworkers.

-
GA20534
Acetyl-Amyloid β-Protein (15-20) amide
Incubation of Ac-QKLVFF-NH? with the amyloid β-protein (1-40) inhibited polymerization of the amyloid β-protein (1-40) into amyloid fibrils. The peptide is thought to block the polymerization sites.

-
GA20533
Acetyl-Amyloid β/A4 Protein Precursor₇₇₀ (96-110) (cyclized)
This cyclized peptide which is homologous to the heparin-binding domain of APP, binds strongly to heparin and inhibits binding of ¹²?I-labeled APP to heparin (IC??= 10??M). The peptide blocks the heparan sulfate proteoglycan-dependent stimulatory effect of APP on neurite outgrowth.

-
GC35334
Amyloid β Peptide (42-1)(human)
Amyloid β Peptide (42-1)(human) is the inactive form of Amyloid β Peptide (1-42).

-
GC35335
Amyloid β-peptide (1-40) rat
Amyloid β-peptide (1-40) rat is a rat form of the amyloid β-peptide, which accumulates as an insoluble extracellular deposit around neurons, giving rise to the senile plaques associated with Alzheimer's disease (AD).

-
GA20721
Amyloid β-Protein (1-12)

-
GA20722
Amyloid β-Protein (1-14)
The N-terminal Aβ fragments Aβ1-14, Aβ1-15 (H-6368), and Aβ1-16 (H-2958) are elevated in cell media and in CSF in response to γ-secretase inhibitor treatment. The presence of these small peptides is consistent with a catabolic amyloid precursor protein cleavage pathway by β- followed by α-secretase. It has been shown that Aβ1-14, Aβ1-15, and Aβ1-16 increase dose-dependently in response to γ-secretase inhibitor treatment while Aβ1-42 levels are unchanged.

-
GA20724
Amyloid β-Protein (1-24)

-
GA20725
Amyloid β-Protein (1-37)
Amyloid β-Protein (1-37) correlates moderately with Mini-Mental State Examination (MMSE) scores in Alzheimer disease.

-
GA20726
Amyloid β-Protein (1-38)
Like Aβ (25-35) (H-1192), the Aβ fragment (1-38) destabilizes calcium homeostasis and renders human cortical neurones vulnerable to environmental insults.

-
GA20727
Amyloid β-Protein (1-39)
Small quantities of Aβ37, 38 and 39 can be detected in CSF together with Aβ40, the most abundant Aβ homolog, Aβ42, and N-terminally truncated amyloid peptides. The relative amounts depend on the variant of Alzheimer's disease. The C-terminally truncated amyloid peptides are also found in amyloid plaques.

-
GA20728
Amyloid β-Protein (1-40) (scrambled)

-
GA20729
Amyloid β-Protein (1-40) amide

-
GA20733
Amyloid β-Protein (1-42)
Compared to the inner salt, the HCl salt of Aβ42 aggregates more readily at pH 7.4.

-
GA20730
Amyloid β-Protein (1-42) (HFIP-treated)
H-7442 was obtained by dissolving Amyloid β-Protein (1-42) (H-1368) in HFIP, aliquoting, and removing the solvent as described in the literature.

-
GA20731
Amyloid β-Protein (1-42) (scrambled)

-
GA20736
Amyloid β-Protein (1-43)
Amyloid β-Protein (1-43) is more prone to aggregation and has higher toxic properties than the long-known Aβ1-42.

-
GA20737
Amyloid β-Protein (1-46)
Precursor of the secreted amyloid β-protein (1-40) and (1-42). The identification of amyloid-β-protein (1-46) led to the identification of a zeta-cleavage site between the known γ- and ε-cleavage sites within the transmembrane domain of amyloid-β precursor protein (APP).

-
GA20738
Amyloid β-Protein (1-6)
Experiments using sub-peptides of Aβ42 revealed that the epitope identified by the antibody A8, as described by Ying and coworkers, lies within the 1-6 region of Aβ. The antibody displays high affinity for soluble Aβ42 oligomers in the molecular weight range of 16.5-25 kDa, and detected target antigen in brain sections from senescence-accelerated SAMP 8 mice.

-
GA20739
Amyloid β-Protein (1-6) amide
Experiments using sub-peptides of Aβ42 revealed that the epitope identified by the antibody A8, as described by Ying and coworkers, lies within the 1-6 region of Aβ. The antibody displays high affinity for soluble Aβ42 oligomers in the molecular weight range of 16.5-25 kDa, and detected target antigen in brain sections from senescence-accelerated SAMP 8 mice. Amidated or acetylated and amidated forms of the sequence were used for example for quantitative structure retention relationships (QSRR) experiments. The latter could allow prediction of reversed-phase high-performance liquid chromatography (HPLC) retention of peptides, as reported by Kaliszan and coworkers.

-
GA20720
Amyloid β-Protein (10-35)
Amyloid β-protein (10-35), YEVHHQKLVFFAEDVGSNKGAIIGLM, was used as a truncated peptide model for the full-length amyloid β-proteins (1-40) and (1-42) in high-resolution structural studies. In contrast to the full-length amyloid β-proteins, amyloid β-protein (10-35) allowed the controlled and reproducible formation of homogeneous fibrils from aqueous solutions of defined pH, ionic strength and soluble peptide concentration necessary for high-resolution structural studies.

-
GA20723
Amyloid β-Protein (11-42)



