Others
- Actin(18)
- Inhibitory Antibodies(137)
- Adenylate cyclase(27)
- AhR(14)
- Aldose reductase(24)
- Arginase(12)
- AT Receptors(1)
- Autotaxin(4)
- ATP citrate lyase (10)
- BCRP(14)
- Caged Compounds(3)
- Calmodulin protein kinase II(1)
- CaM kinase II(10)
- Carbonic Anhydrase(21)
- Cell Penetrating Peptide(2)
- Ceramidases(4)
- CERK(1)
- Collagen proline hydroxylase(1)
- COT/TpI2(0)
- COMT(12)
- CPA(0)
- CPG2(0)
- Cytokine(4)
- dCTP Pyrophosphatase(1)
- DHODH(1)
- Diacylglycerol Kinase(1)
- DNA Stain(16)
- eIF2a(2)
- Endopeptidase(3)
- Enzyme Substrates / Activators(12)
- ES-FLI1/RHA(2)
- Farnesyl Diphosphate Synthase(1)
- FBPase(4)
- FLAP(9)
- Fluorescent Probes(232)
- GLI(3)
- Glycoprotein(5)
- Glycosylase(15)
- Saccharides and Glycosides(13)
- Glyoxalase I(1)
- GSNOR(2)
- GSTP1-1(0)
- Guanylate Cyclase (27)
- Guanylyl Cyclase(7)
- Gutathione S-transferase(12)
- Heme oxygenase(7)
- Hexokinase(9)
- Hydroxylases(10)
- IL Receptor(1)
- Imidazoline Receptors(17)
- IMPDH(1)
- Inositol Phosphatases(4)
- Lp-PLA2(1)
- LpxC(1)
- Matrix Metalloprotease(1)
- MDR multidrug resistance(35)
- Menin-MLL(5)
- MGMT(1)
- Mineralocorticoid Receptor(20)
- Miscellaneous Compounds(9)
- MyD88(7)
- Myosin(23)
- NAE(2)
- NADPH Oxidase(14)
- Nampt(21)
- Natriuretic Peptide Receptors(4)
- Nitric Oxide(62)
- Nuclear Receptors(13)
- Oxygenases/Oxidases(5)
- PDHK(9)
- PGC-1α(4)
- Pregnane X Receptors(2)
- Reductases(7)
- Renin(14)
- Retinoid X Receptors(11)
- Squalene epoxidase(4)
- SGK(9)
- sFRP-1(1)
- SPHK(18)
- Synthases/Synthetases(12)
- Tachykinin(8)
- Thrombopoietin Receptor(7)
- Thymidine phosphorylase(4)
- Transcription Factors(84)
- Transketolase(3)
- β(1,3)-D-Glucan Synthase(2)
- β-Lactamase(3)
- XAO(4)
- Others(1640)
- BMI-1(1)
- Hydrogen Peroxide(0)
- Ascorbic Acid(0)
- SOD(3)
- Homodimerizers(2)
- Renal Diseases(9)
- PAF-AH(1)
- FAS(30)
- calmodulin(7)
- Protein synthesis(5)
- Epac(1)
- MAGL(20)
- exocytosis(1)
- peroxiredoxins(1)
- Glucosidase(49)
- Estrogen Receptor/ERR(164)
- Progesterone Receptor(35)
- Cyclase(5)
- Environmental Toxicology(164)
- Glycerolipid Lipases(12)
- Lipases(23)
- Multidrug-Resistance Proteins(8)
- Negative Controls(8)
- Nucleic Acid Turnover/Signaling(34)
- Organ- & System-Specific Toxicity(105)
- Pesticides(57)
- Pharmaceutical Impurities(24)
- Sulfotransferases(3)
- Tec(1)
- Toxicology(104)
- Xenobiotic Sensing(18)
- UDP Glucuronosyltransferases(38)
Products for Others
- Cat.No. Product Name Information
-
GC19789
Hyaluronidase
Hyaluronidase;>300u/mg
-
GC45195
α,β-Methyleneadenosine 5'-triphosphate (sodium salt)
αβ-methylene ATP
α,β-Methyleneadenosine 5'-triphosphate (sodium salt), a phosphonic analog of ATP, is a P2X3 and P2X7 receptor ligand.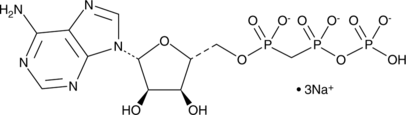
-
GC63268
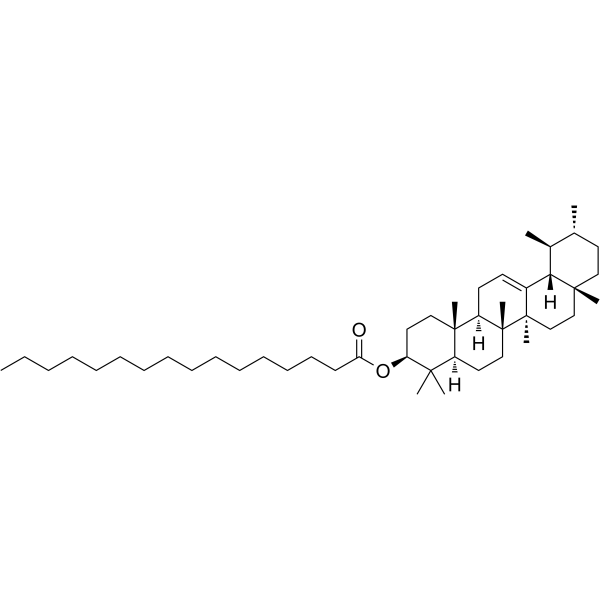
α-Amyrin palmitate
α-Amyrin palmitate is isolated from Santalum album (sandalwood).
-
GC38042
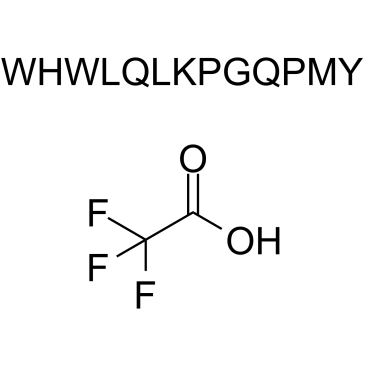
α-Factor Mating Pheromone, yeast (TFA)
Mating Factor α TFA
-
GC45208
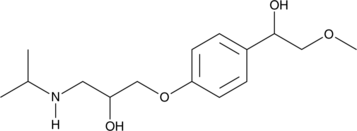
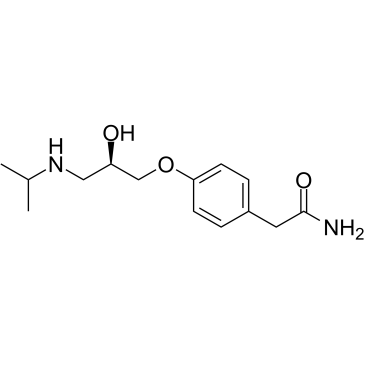
α-hydroxy Metoprolol
α-hydroxy Metoprolol is an active metabolite of the β1-adrenergic receptor blocker metoprolol.
-
GC40302
α-hydroxy Tamoxifen
(E)-α-Hydroxy tamoxifen; α-OHTAM
Tamoxifen is a selective estrogen receptor (ER) modulator that is widely used in the therapeutic and chemopreventive treatment of breast cancer.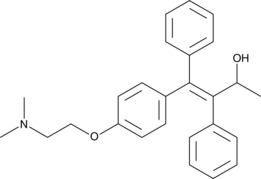
-
GC67685
α-Isopropylmalate
α-IPM

-
GC45210
α-Linolenic Acid (sodium salt)
ALA, C18:3 (9Z,12Z,15Z), C18:3 n-3
α-Linolenic acid (ALA) is an essential fatty acid found in leafy green vegetables.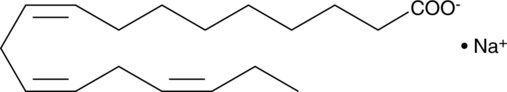
-
GC48283
α-Linolenic Acid-d14
ALAd14
An internal standard for the quantification of αLinolenic acid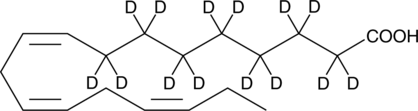
-
GC45602
α-Linolenic Acid-d5 MaxSpec• Standard
ALA-d5, C18:3 (9Z,12Z,15Z)-d5, C18:3 n-3-d5

-
GC65892
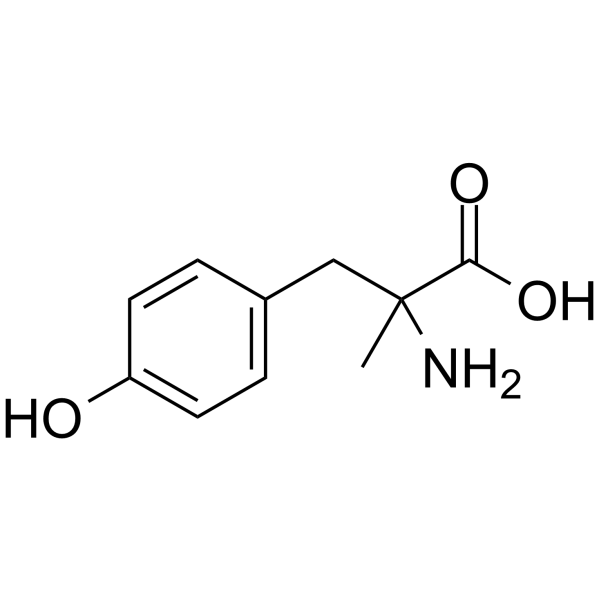
α-Methyl-p-tyrosine
α-Methyl-p-tyrosine is a competitive inhibitor of the enzyme tyrosine hydroxylase, which converts tyrosine to Levodopa (DOPA). α-Methyl-p-tyrosine is an orally active inhibitor of catecholamine synthesis which inhibits the hydroxylation of tyrosine to DOPA.
-
GC37980
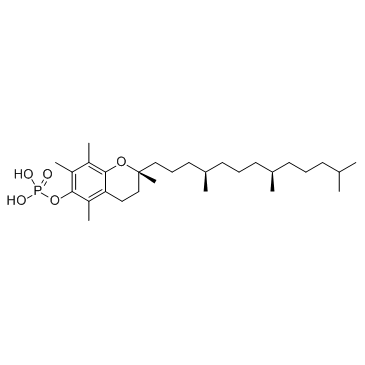
α-Tocopherol phosphate
α-Tocopherol phosphate is the compound demonstrating the highest vitamin E activity, which is available both in its natural form as RRR-alpha-tocopherol isolated from plant sources.
-
GC45224
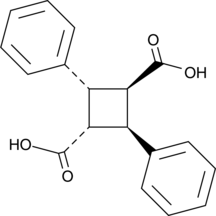
α-Truxillic Acid
Gratissimic Acid
α-Truxillic acid can be formed by the dimerization of two molecules of α-trans-cinnamic acid.
-
GC45603
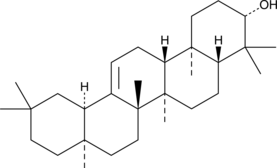
β-Amyrin
(+)-β-Amyrin, NSC 527971, Olean-12-en3β-ol
-
GC41267
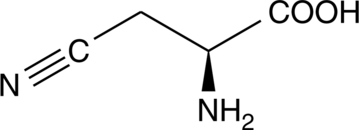
β-cyano-L-Alanine
BCA
Hydrogen sulfide (H2S) is a naturally-occurring gasotransmitter with vasodilator and inflammatory modulating activity.
-
GC63276
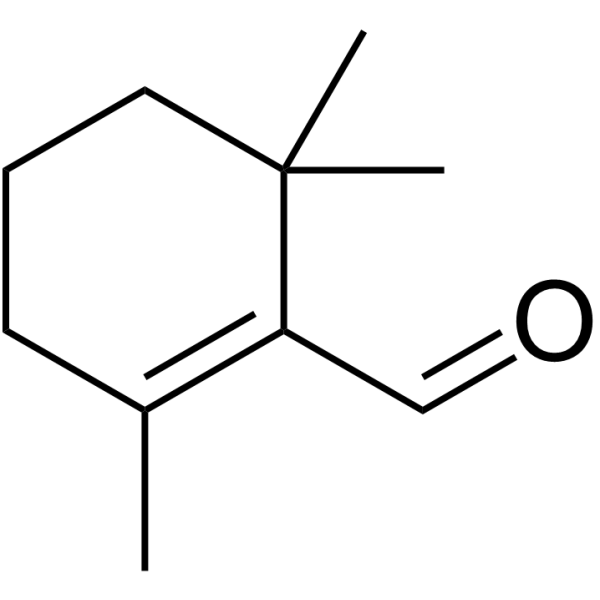
β-Cyclocitral
β-Cyclocitral, a volatile oxidized derivative of β-carotene, is a grazer defence signal unique to the Cyanobacterium Microcystis.
-
GC63277
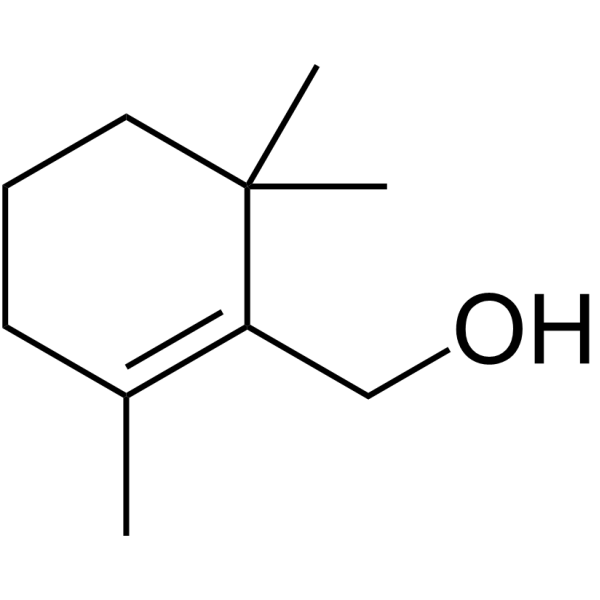
β-Cyclogeraniol
β-Cyclogeraniol is a natural odour compound.
-
GA24007
β-Endorphin (30-31) (bovine, camel, mouse, ovine)
β-Endorphin (30-31) (bovine, camel, mouse, ovine) (Glycyl-L-glutamine), as a enzymatic cleavage product of β-endorphin, is apparently an endogenous antagonist of beta-endorphin(1-31) in several systems.

-
GC70186
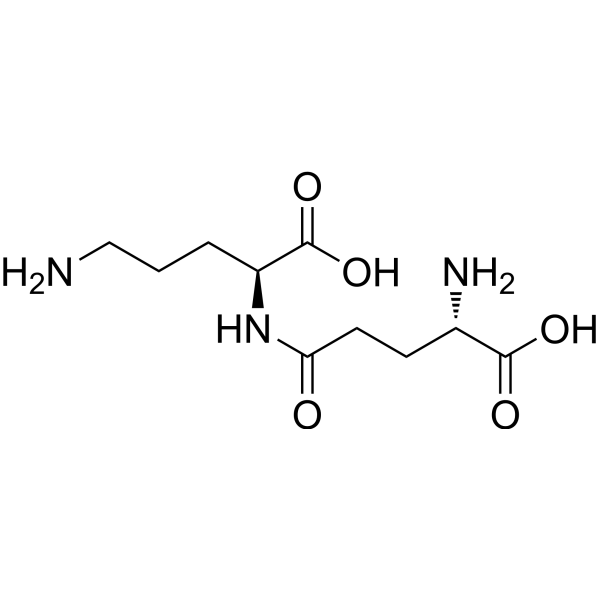
γ-Glutamylornithine
Gamma-glutamylornithine is a urine excretion product of patients with HHH syndrome (hyperuricemia, hyperammonemia, and homocitrullinuria) and patients with uric acid-related recurrent ataxia. Increased endogenous ornithine levels lead to an increase in gamma-glutamylornithine levels in the urine.
-
GC63280
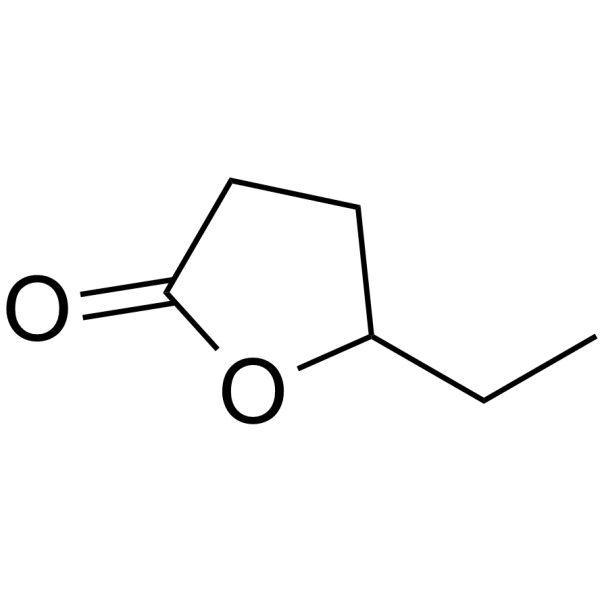
γ-Hexalactone
γ-Hexalactone is a gamma-lactone found in ripe fruits.
-
GC31977
α-2,3-sialyltransferase-IN-1
Lith-O-Asp analog
α-2,3-sialyltransferase-IN-1 (Lith-O-Asp analog) is a noncompetitive α-2,3-sialyltransferase inhibitor with an IC50 of 6 μM.
-
GC30203
α-Amylase

-
GC30587
α-Factor Mating Pheromone, yeast (Mating Factor α)
α-Factor Mating Pheromone, yeast (Mating Factor α) is a tridecapeptide secreted by S.
-
GC33470
β-Apo-13-carotenone (D'Orenone)
D'Orenone
β-Apo-13-carotenone (D'Orenone) (D'Orenone) is a naturally occurring β-apocarotenoid functioned as an antagonist of RXRα.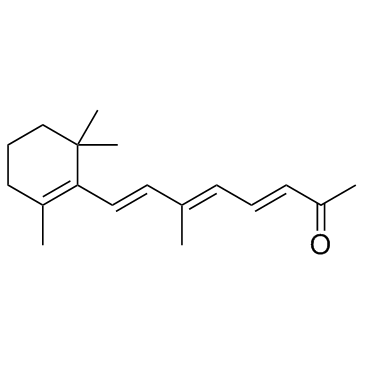
-
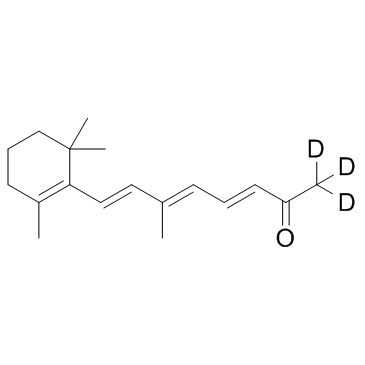
GC33505
β-Apo-13-carotenone D3 (D'Orenone D3)
-
GC30207
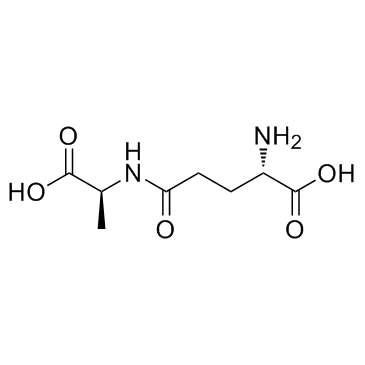
γ-L-Glutamyl-L-alanine
γ-L-Glutamyl-L-alanine, composed of gamma-glutamate and alanine, is a proteolytic breakdown product of larger proteins.
-
GC63281
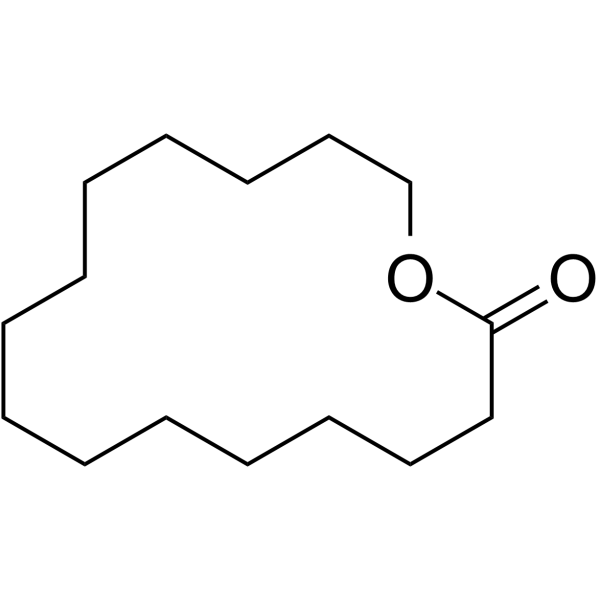
ω-Pentadecalactone
ω-Pentadecalactone is a fragrance ingredient.
-
GC70556
ß-pBrPh-Glc
β-pBrPh-Glc is a small-molecule ice recrystallization inhibitor.

-
GC16005
(±)-CPSI 1306
macrophage inhibitory factor (MIF) inhibitor

-
GC46289
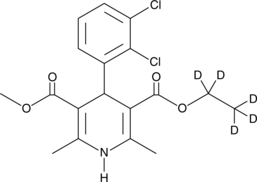
(±)-Felodipine-d5
An internal standard for the quantification of (±)-felodipine
-
GC40843
(±)-Goitrin
DL-Goitrin, (R,S)-Goitrin
(±)-Goitrin, also called (R, S)- report by the spring, consists of the epigoitrin (reported by the R- Spring) and the spring (-S- reported by spring), and the two mutually isomers, and the mixture is the ingredient of Radix.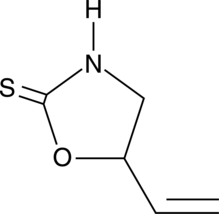
-
GC41672
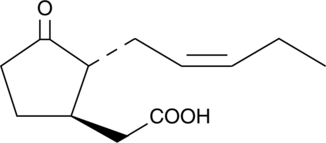
(±)-Jasmonic Acid
Jasmonic acid, derived from α-linolenic acid , is a plant growth regulator involved in the signaling mechanisms for a variety of conditions including plant defense, wound healing, tuberization, fruit ripening, and senescence.
-
GC41677
(±)-SDZ-201 106
DPI 201-106
(±)-SDZ-201 106 (SDZ 201106) is a cardiotonic agent with a synergistic sarcolemmal and intracellular mechanism of action.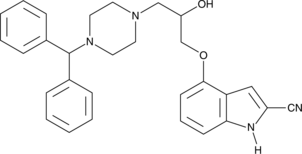
-
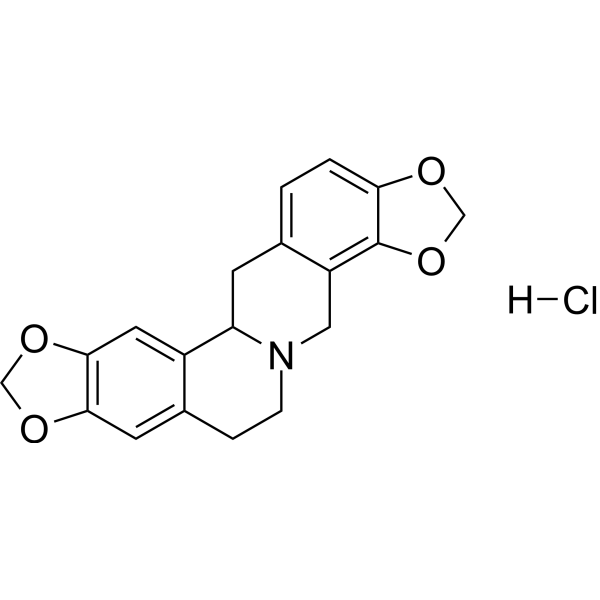
GC67928
(±)-Stylopine hydrochloride
Tetrahydrocoptisine hydrochloride
-
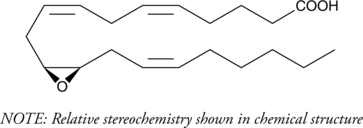
GC40466
(±)11(12)-EET
(±)11,12-EpETrE
(±)11(12)-EET is a fully racemic version of the R/S enantiomeric forms biosynthesized from arachidonic acid by cytochrome P450 enzymes.
-
GC40802
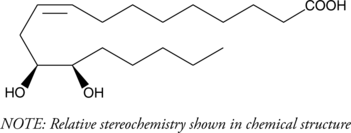
(±)12(13)-DiHOME
Isoleukotoxin diol
(±)12(13)-DiHOME is the diol form of (±)12(13)-EpOME, a cytochrome P450-derived epoxide of linoleic acid also known as isoleukotoxin.
-
GC40434
(±)16-HETE
(±)16-Hydroxyeicosatetraenoic Acid
Electrolyte and fluid transport in the kidney are regulated in part by arachidonic acid and its metabolites.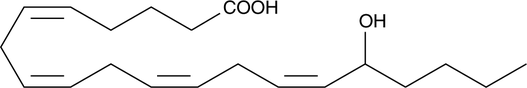
-
GC40435
(±)17-HETE
(±)-Hydroxyeicosatetraenoic Acid
Electrolyte and fluid transport in the kidney are regulated in part by arachidonic acid and its metabolites.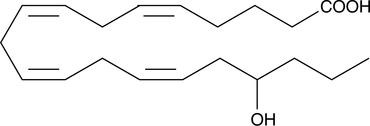
-
GC40436
(±)18-HETE
(±)18-Hydroxyeicosatetraenoic Acid
(±)18-HETE is the racemic version of a cytochrome P450 (CYP450) metabolite of arachidonic acid.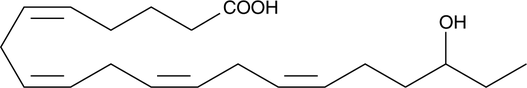
-
GC40801
(±)9(10)-DiHOME
Leukotoxin diol
Leukotoxin is the 9(10) epoxide of linoleic acid, generated by neutrophils during the oxidative burst.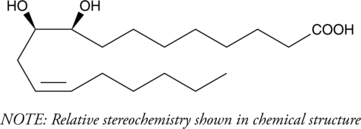
-
GC30358
(±)-1,2-Propanediol (1,2-(RS)-Propanediol)
(±)-1,2-Propanediol (1,2-(RS)-Propanediol) (1,2-(RS)-Propanediol) is an aliphatic alcohol and frequently used as an excipient in many drug formulations to increase the solubility and stability of drugs.

-
GC32617
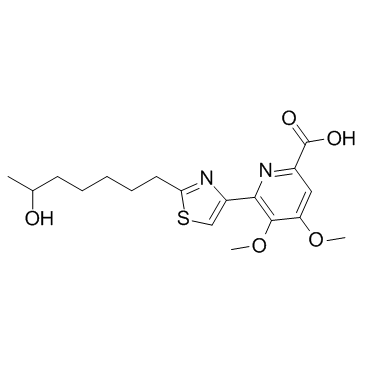
(±)-WS75624B
(±)-WS75624B is an endothelin converting enzyme (ECE) inhibitor with an IC50 of 0.03 μg/mL.
-
GC34953
(+)-BAY-1251152
(+)-BAY-1251152; (+)-VIP152
(+)-BAY-1251152 ((+)-BAY-1251152) is an enanthiomer of BAY-1251152 with rotation (+). (+)-BAY-1251152 is a potent and selective CDK9 inhibitor with an IC50 of 3 nM. (+)-BAY-1251152 has anti-tumour activity.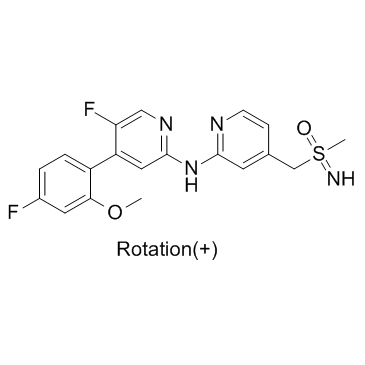
-
GC63999
(+)-Coclaurine hydrochloride
(+)-(R)-Coclaurine hydrochloride; (R)-Coclaurine hydrochloride; d-Coclaurine hydrochloride
(+)-Coclaurine ((+)-(R)-Coclaurine) hydrochloride, benzyltetrahydroisoquinoline alkaloid isolated from a variety of plant sources.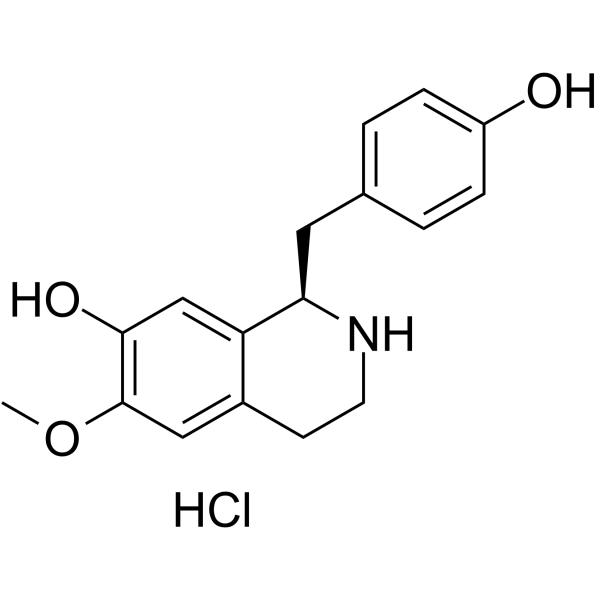
-
GC38119
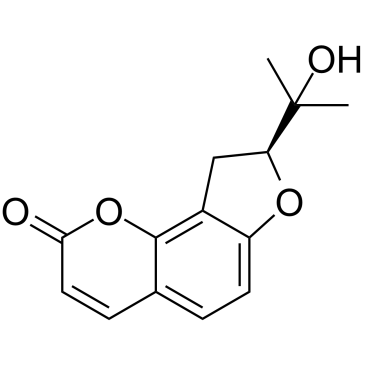
(+)-Columbianetin
(+)-Columbianetin is an isomer of Columbianetin.
-
GC70265
(+)-EMD 57033
(+)-EMD 57033 is a cardiac troponin C (cTnC) activator, is a dominant Ca2+ sensitizer.

-
GC61765
(+)-Isomenthone
(±)-Isomenthone, cis-Menthone
(+)-Isomenthone is an isomenthone isolated from Ziziphora clinopodioides Lam.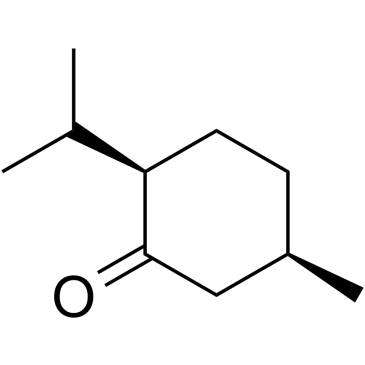
-
GC38443
(+)-SHIN1
RZ2994
(+)-SHIN1 ((+)-RZ-2994) is an active (+) enantiomer of SHIN1.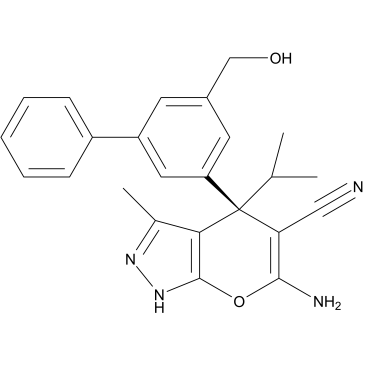
-
GC17330
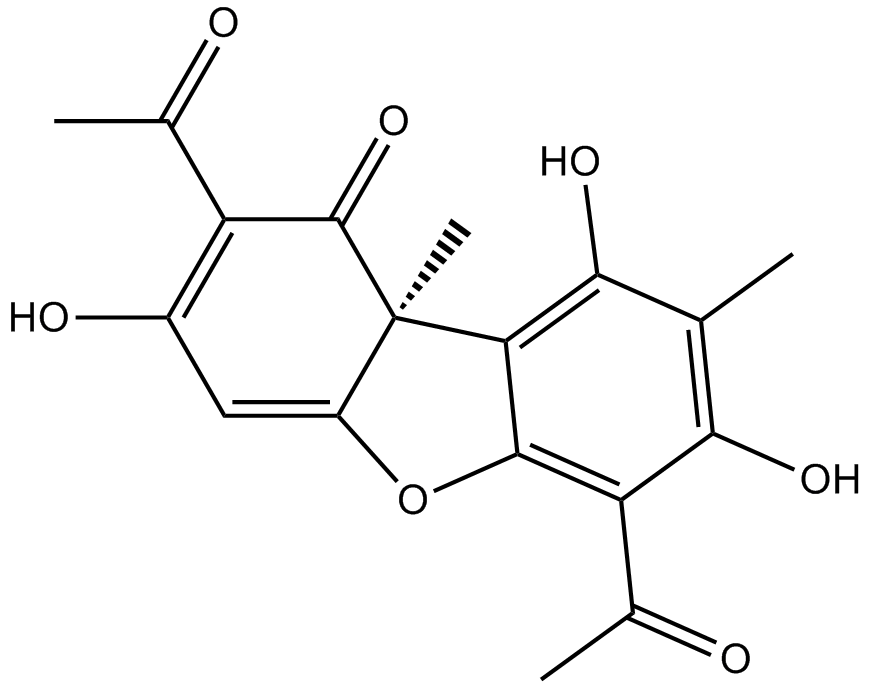
(+)-Usniacin
(+)-Usniacin is isolated from isolated from lichens, binds at the ATP-binding pocket of mTOR, and inhibits mTORC1/2 activity. (+)-Usniacin inhibits the phosphorylation of mTOR downstream effectors: Akt (Ser473), 4EBP1, S6K, induces autophay, with anti-cancer activity. (+)-Usniacin possesses antimicrobial activity against a number of planktonic gram-positive bacteria, including Staphylococcus aureus, Enterococcus faecalis, and Enterococcus faecium.
-
GC69720
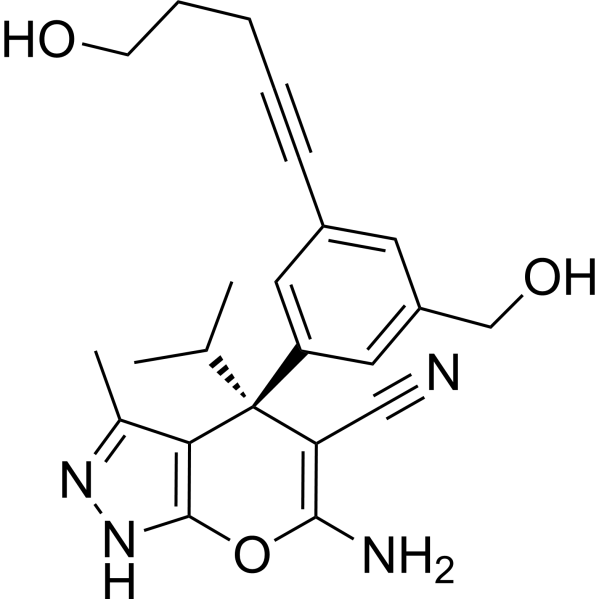
(+)SHIN2
(+)SHIN2 is a serine hydroxymethyltransferase (SHMT) inhibitor, and its in vivo target can be traced using 13C-serine. (+)SHIN2 increases the survival rate of mice with primary acute T-cell lymphoblastic leukemia (T-ALL) driven by Notch1 and has a synergistic effect with Methotrexate.
-
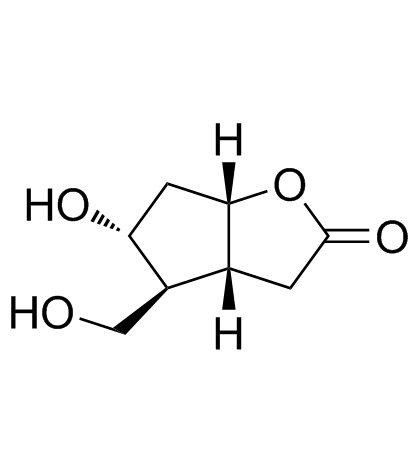
GC30750
(-)-Corey lactone diol
-
GC34948
(-)-GSK598809
1S,5R-GSK598809
(-)-GSK598809 is an isomer of GSK598809.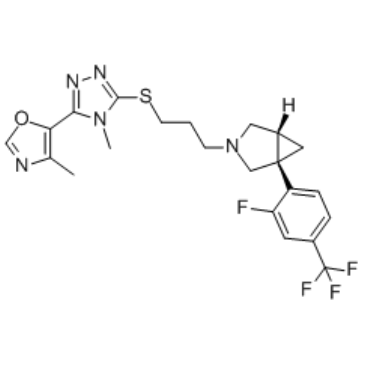
-
GC41344
(-)-Guaiol
NSC 19941
(-)-Guaiol is a sesquiterpene alcohol that has been found in several traditional Chinese medicinal plants and has antiproliferative, pro-autophagic, insect repellent, and insecticidal biological activities.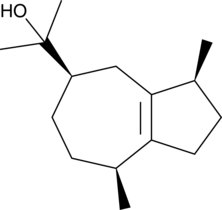
-
GC14520
(-)-p-Bromotetramisole Oxalate
(-)-p-Bromotetramisole, L-para-Bromotetramisole
(-)-p-Bromotetramisole Oxalate(L-p-bromotetramisole) is a potent and non-specific inhibitor of alkaline phosphatase and is also an inhibitor of protein tyrosine phosphatases.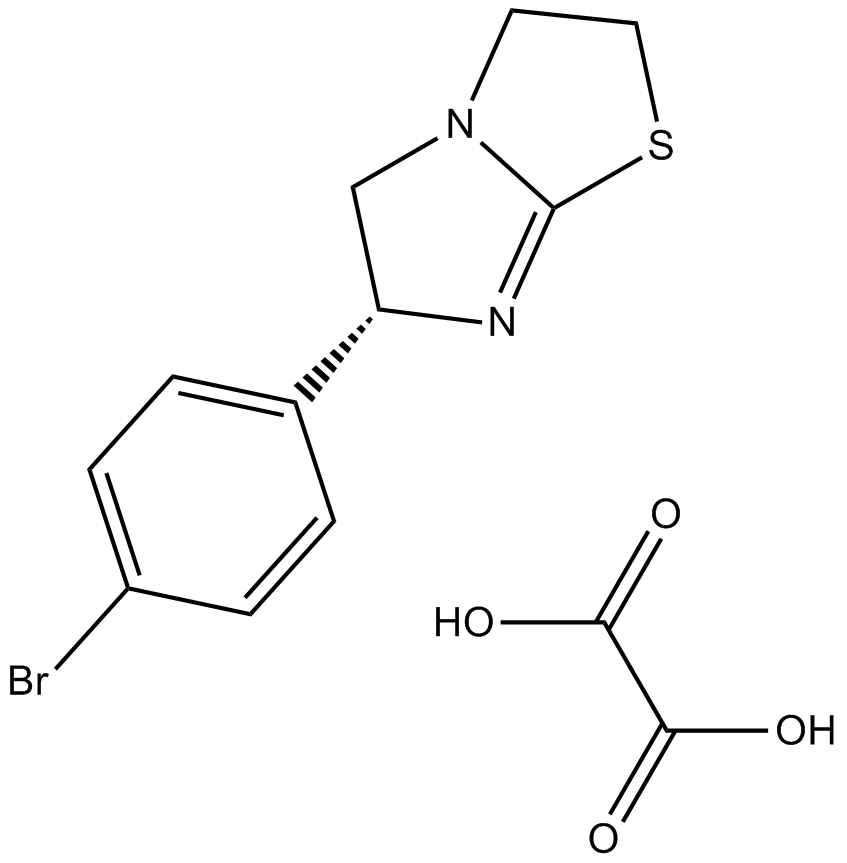
-
GC38444
(-)-SHIN1
RZ2994
(-)-SHIN1 ((-)-RZ-2994) is an inactive (?) enantiomer of SHIN1.
-
GC12164
(-)-Terreic acid
TA, (-)-Terreic Acid
Bruton's tyrosine kinase (BTK) inhibitor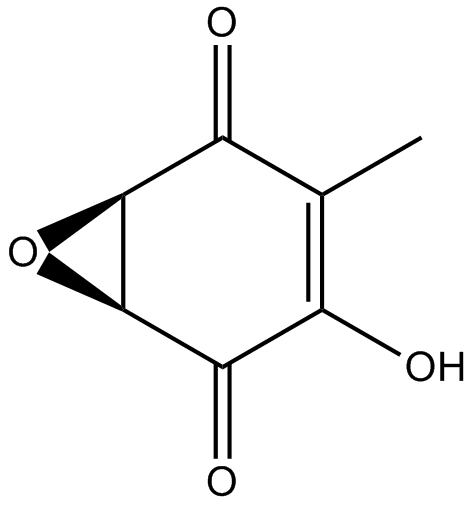
-
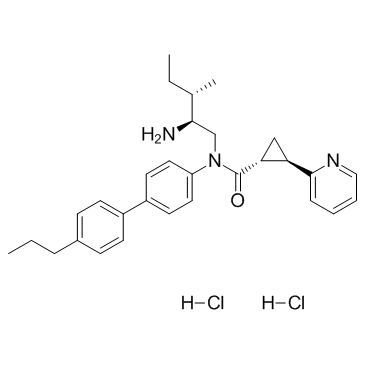
GC30855
(1R,2R)-2-PCCA(hydrochloride)
(1R,2R)-2-PCCA(hydrochloride) is a diastereomer of 2-PCCA, and acts as a potent GPR88 receptor agonist, with an EC50 of 3 nM in cell-free assay, and 603 nM in cell assay.
-
GC67880
(1R,2S)-Xeruborbactam disodium
(1R,2S)-QPX7728 disodium

-
GC62730
(1S)-Calcitriol
1α,25-Dihydroxy-3-epi-vitamin-D3
(1S)-Calcitriol (1α,25-Dihydroxy-3-epi-vitamin-D3) is a natural metabolite of 1α,25-dihydroxyvitamin D3 (1α,25(OH)2D3).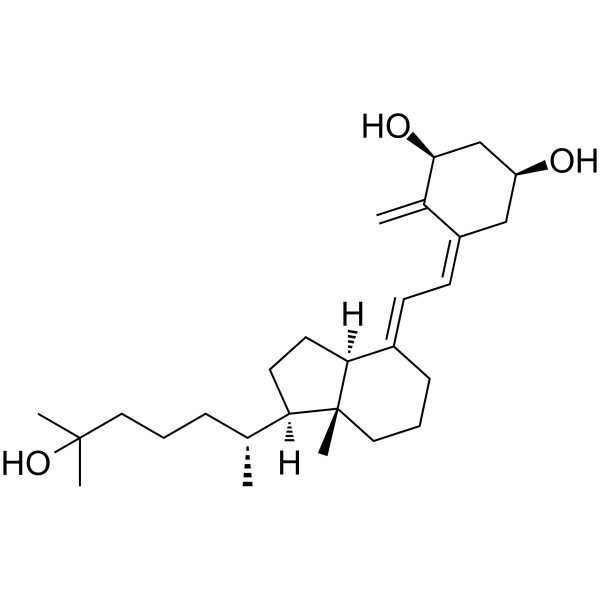
-
GC34440
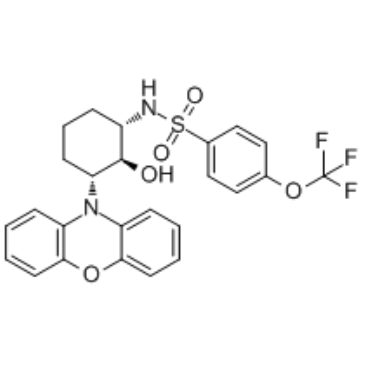
(1S,2S,3R)-DT-061
(1S,2S,3R)-DT-061 is an enantiomer of DT-061. DT-061 is an orally bioavailable activator of protein phosphatase 2A (PP2A) and could be applied in the therapy of KRAS-mutant and MYC-driven tumorigenesis.
-
GC70234
(2R)-Octyl-α-hydroxyglutarate sodium
(2R)-Octyl-α-droxyglutarate (sodium) is the sodium salt form of (2R)-Octyl-α-droxyglutarate.

-
GC32754
(2R)-Octyl-α-hydroxyglutarate ((2R)-Octyl-2-HG)
(2R)Octyl2-HG
(2R)-Octyl-α-hydroxyglutarate ((2R)-Octyl-2-HG) ((2R)-Octyl-2-HG) is a modified form of D-isomer 2-Hydroxyglutarate.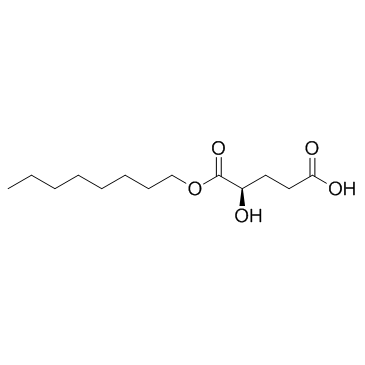
-
GC70210
(2R)-SR59230A
(2R)-SR59230A is the isomer of SR59230A , and can be used as an experimental control.

-
GC68522
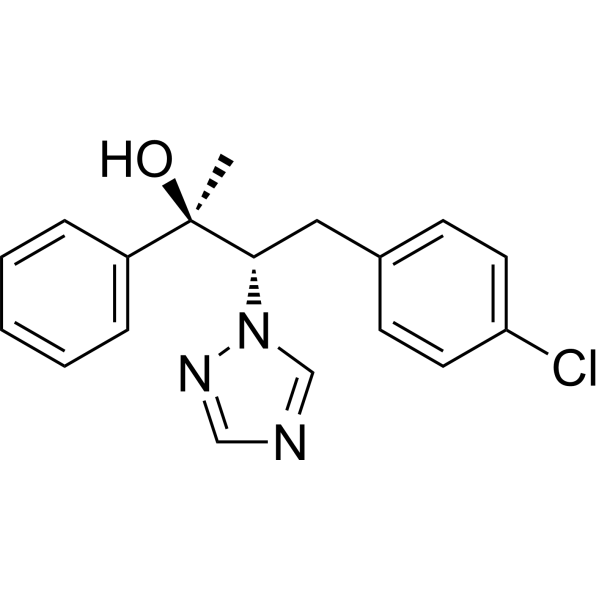
(2R,3S)-Brassinazole
Brassinazole (0.5, 1, 5 μM) significantly caused morphological abnormalities in seedlings similar to BR-deficient mutants. Brassinazole resulted in dwarfism and altered leaf morphology, such as the typical downward curling and dark green appearance seen in Arabidopsis BR-deficient mutants. However, treatment with 10 nM BR reversed the dwarfism.
-
GC34970
(2S)-2'-Methoxykurarinone
2'-O-Methylkurarinone
(2S)-2'-Methoxykurarinone, a compound isolated from the roots of Sophora flavescens, has anti-inflammatory, antipyretic, antidiabetic, and antineoplastic effects.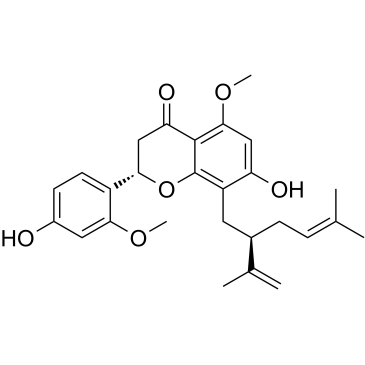
-
GC19473
(2S)-Octyl-α-hydroxyglutarate
(2S)-Octyl-2-HG
A modified form of S-isomer 2-Hydroxyglutarate

-
GC68523
(2S,3R)-Brassinazole
(2S,3R)-Brassinazole is an isomer of brassinazole, which is a compound that inhibits the biosynthesis of brassinosteroids (BR), a class of plant steroids, by acting on the oxidation process from 6-oxo-campestanol to teasterone. (2S,3R)-Brassinazole may be the most active form of brassinazole.

-
GC38874
(2S,3R,4S)-4-Hydroxyisoleucine
(4S)-4-Hydroxy-L-isoleucine, 4-OH-Ile
(2S,3R,4S)-4-Hydroxyisoleucine is an orally active compound isolated from Trigonella foenum-graecum, with anti-diabetes and anti-diabetic nephropathy activity.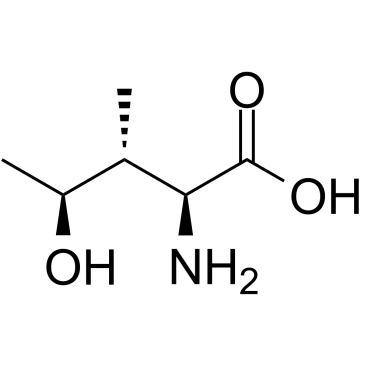
-
GC30605
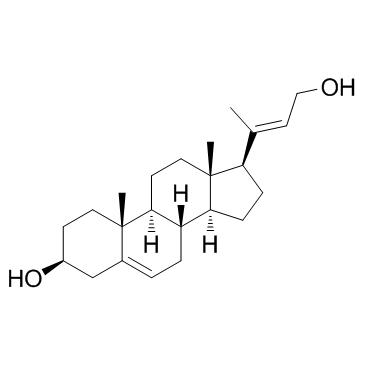
(3β,20E)-24-Norchola-5,20(22)-diene-3,23-diol
(3β,20E)-24-Norchola-5,20(22)-diene-3,23-diol is a steroid-based allylic alcohol.
-
GC72663
(3E,8Z,11Z)-3,8,11-Tetradecatrienyl acetate
(3E,8Z,11Z)-3,8,11-Tetradecatrienyl acetate is a sex pheromone capable of attracting male South American tomato pinworms (Scrobipalpuloides absoluta), which can be isolated from the tomato pest.

-
GC13782
(3S,4S)-3-(Boc-amino)-4-methylpyrrolidine
(3S,4S)-3-(Boc-amino)-4-methylpyrrolidine

-
GC65358
(4-NH2)-Exatecan
(4-NH2)-Exatecan, a topoisomerase inhibitor derivative extracted from patent US20200306243A1, compound A. (4-NH2)-Exatecan can be used in the synthesis of antibody-drug conjugates (ADCs).

-
GC39320
(5E,9E,13E)-Teprenone
(5E,9E,13E)-Geranylgeranylacetone
(5E,9E,13E)-Teprenone ((5E,9E,13E)-Geranylgeranylacetone) is an isomer of Teprenone with antiulcer activity.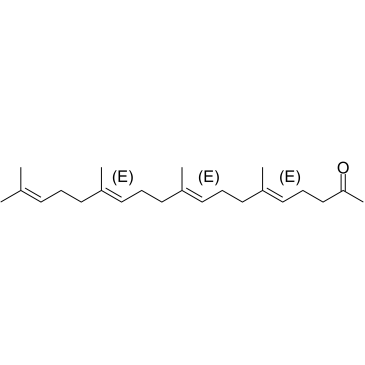
-
GC30701
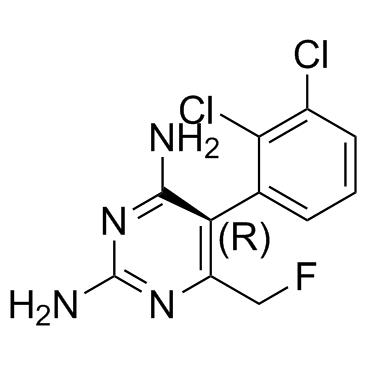
(5R)-BW-4030W92
(5R)-BW-4030W92 is the R enantiomer of BW-4030W92.
-
GC34974
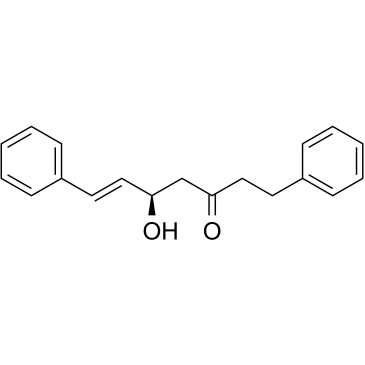
(5R,6E)-5-Hydroxy-1,7-diphenyl-6-hepten-3-one
(5R,6E)-5-Hydroxy-1,7-diphenyl-6-hepten-3-one is the methylene chloride extract of Alpinia nutans, has antioxidant activity.
-
GC13944
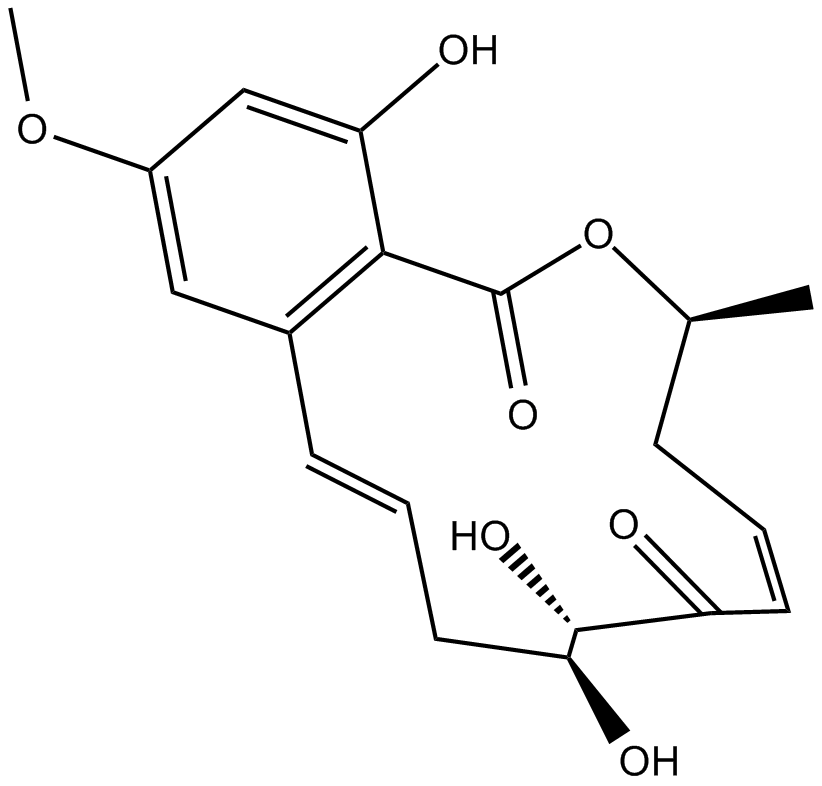
(5Z)-7-Oxozeaenol
FR148083,L-783,279,LL-Z 1640-2
TAK1 mitogen-activated protein kinase kinase kinase (MAPKKK) inhibitor
-
GC15970
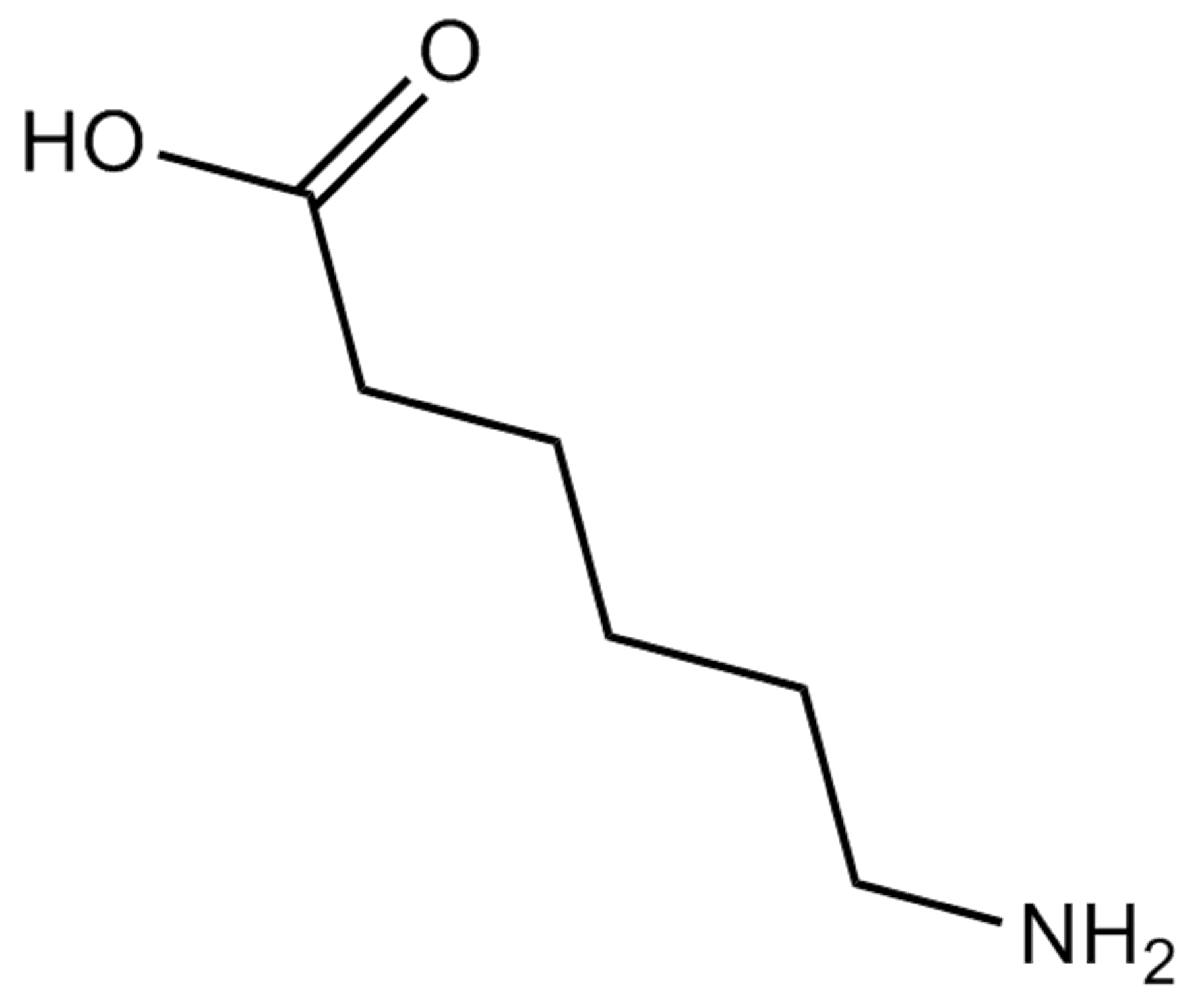
(6-)ε-Aminocaproic acid
εAminocaproic Acid, EACA, NSC 26154, NSC 212532, NSC 400230
antifibrinolytic agent
-
GC68574
(6R)-ML753286
(6R)-ML753286 is an isomer of ML753286. ML753286 is an orally active and selective inhibitor of BCRP (breast cancer resistance protein), with an IC50 of 0.6 μM. ML753286 has high permeability and low to moderate clearance in rodent and human liver S9 fractions, and plasma stability across different species.

-
GC63685
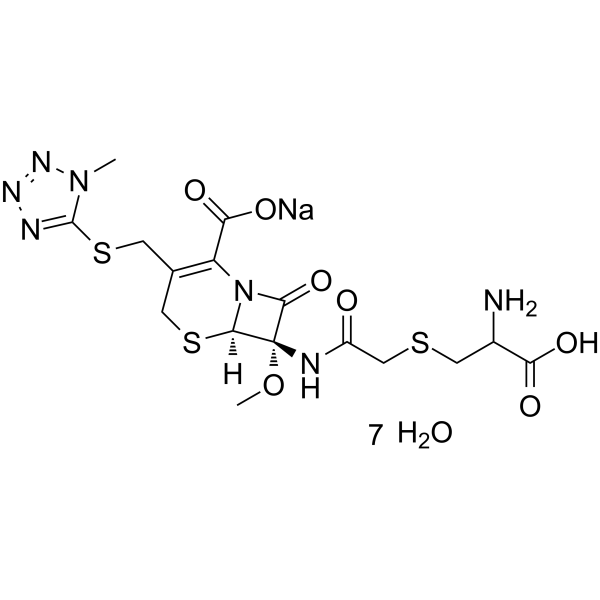
(6R,7S)-Cefminox sodium heptahydrate
(6R,7S)-Cefminox sodium heptahydrate is an isomer of Cefminox sodium heptahydrate.
-
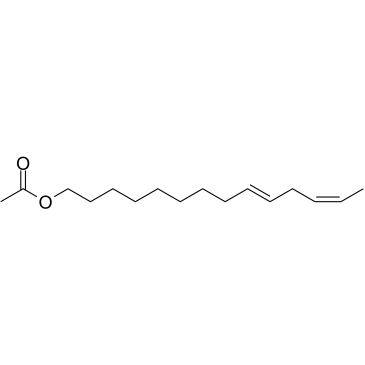
GC34975
(9Z,12E)-Tetradecadien-1-yl acetate
-
GC34976
(Arg)9
(Arg)9 (Nona-L-arginine;Peptide R9) is a cell-penetrating peptide; exhibits neuroprotective activity with an IC50 of 0.78 μM in the glutamic acid model.

-
GC15373
(E)-2-Decenoic acid
trans-2-Decylenic Acid
An unsaturated fatty acid found in royal jelly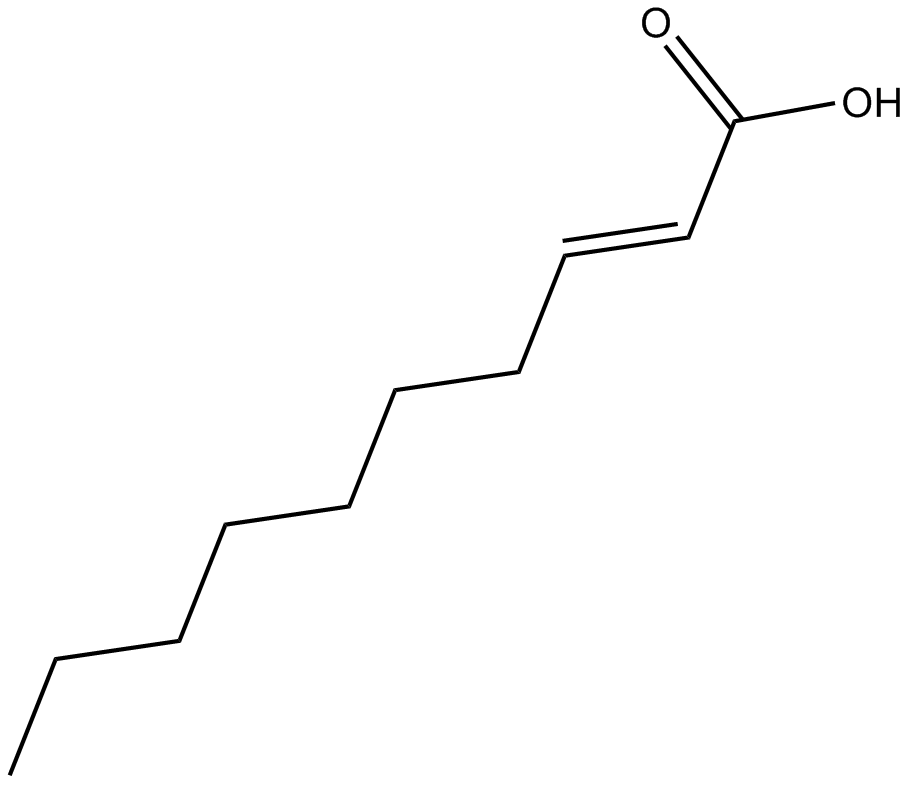
-
GC32522
(E)-Alprenoxime (CDDD-1815)
CDDD-1815
(E)-Alprenoxime (CDDD-1815) is the isomer of the Alprenoxime.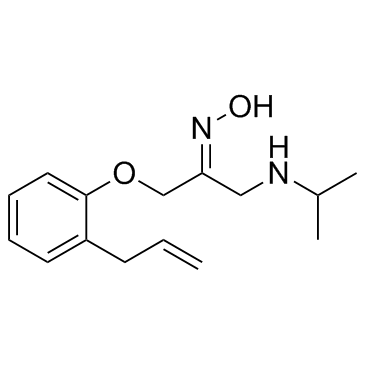
-
GC70220
(E)-Crotylbarbital
(E)-Crotylbarbital is the isomer of Crotylbarbital.

-
GC34982
(E)-LHF-535
(E)-LHF-535 is the E-isomer of LHF-535.

-
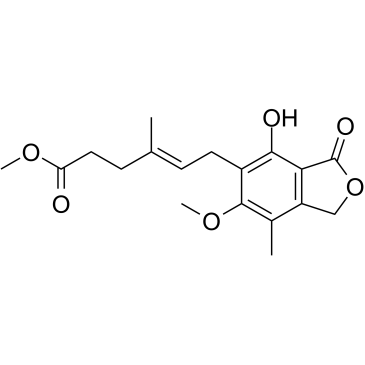
GC60403
(E/Z)-Methyl mycophenolate
(E/Z)-Methyl mycophenolate is a racemic compound of (Z)-Methyl mycophenolate and (E)-Methyl mycophenolate isomers.
-
GC38229
(R)-(+)-Atenolol
An enantiomer of (±)-atenolol
-
GC61425
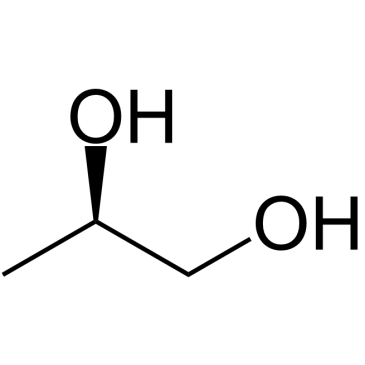
(R)-(-)-1,2-Propanediol
(R)-(-)-1,2-Propanediol is a (R)-enantiomer of 1,2-Propanediol that produced from glucose in Escherichia coli expressing NADH-linked glycerol dehydrogenase genes.
-
GC61694
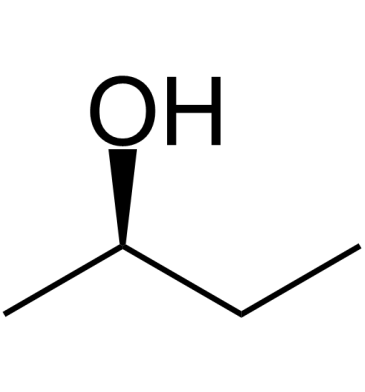
(R)-(-)-2-Butanol
(R)-(-)-2-Butanol is released by the females of the white grub beetle, Dasylepida ishigakiensis, to attract males.
-
GC13030
(R)-(-)-Ibuprofen
(-)-Ibuprofen
Inhibitor of Cox-1 and Cox-2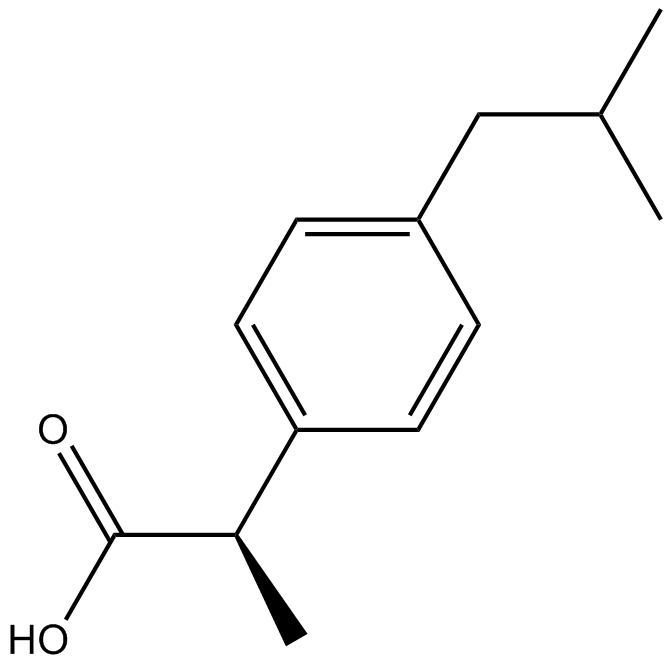
-
GC33574
(R)-1,2,3,4-Tetrahydro-3-isoquinolinecarboxylic acid
(R)-1,2,3,4-Tetrahydro-3-isoquinolinecarboxylic acid is a constrained Phe analogue which can fold into a beta-bend and a helical structure, and to adopt a preferred side-chain disposition in the peptide.

-
GC38716
(R)-Apremilast
(R)-Apremilast ((R)-CC-10004) is a enantiomer of Apremilast.

-
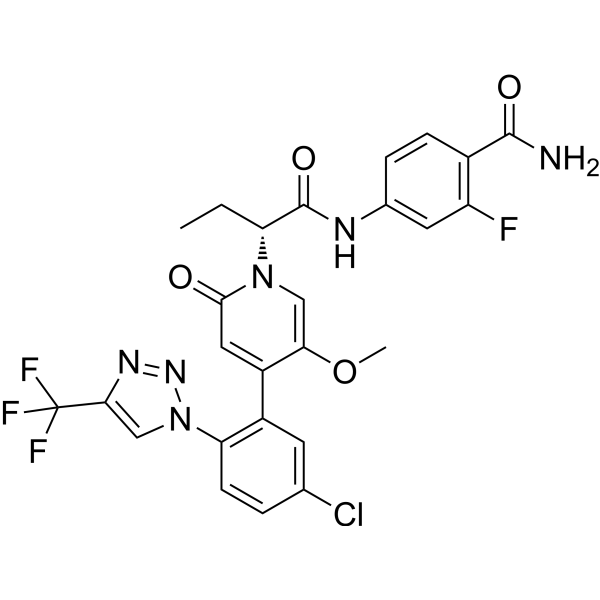
GC67688
(R)-Asundexian
(R)-BAY-2433334
-
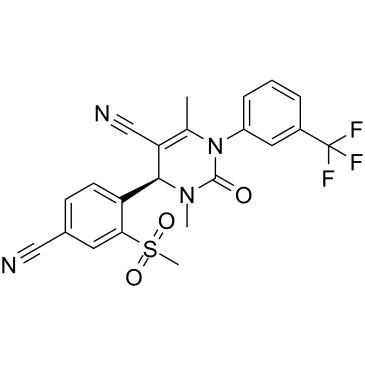
GC34987
(R)-BAY-85-8501
(R)-BAY-85-8501 is the less active Enantiomer of BAY-85-8501.
-
GC63781
(R)-BAY-899
(R)-BAY-899 is the R-enantiomer of BAY-899.

-
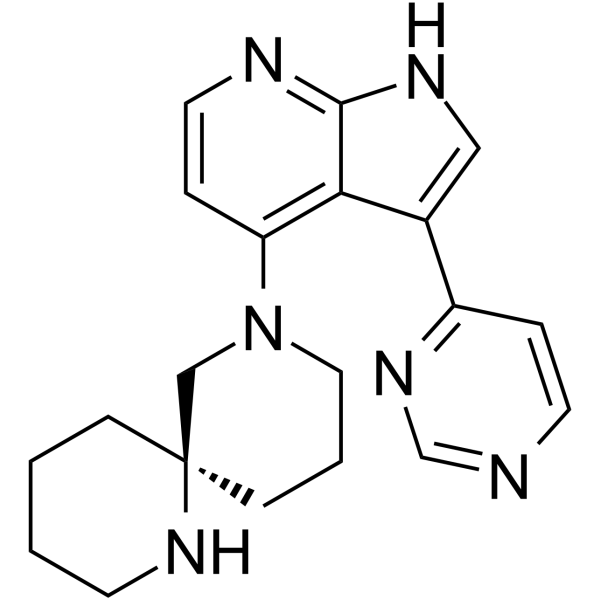
GC63906
(R)-BDP9066
(R)-BDP9066 is a potent inhibitor of myotonic dystrophy kinase-related Cdc42-binding kinase (MRCK). (R)-BDP9066 blocks cancer cell invasion. (R)-BDP9066 has the potential for the research of proliferative diseases, such as cancer.
-
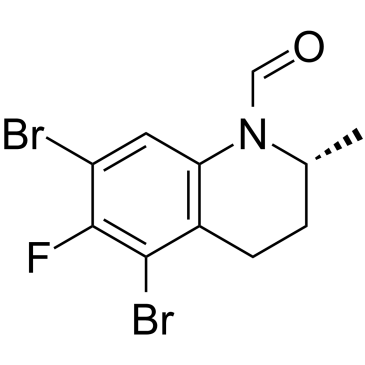
GC34988
(R)-CE3F4
(R)-CE3F4 is a potent and selective inhibitor of exchange protein directly activated by cAMP isoform 1 (Epac1), with an IC50 of 4.2 μM, with 10-fold selectivity for Epac1 over Epac2 (IC50, 44 μM).
-
GC38717
(R)-CSN5i-3
(R)-CSN5i-3 is the (R)-enantiomer of CSN5i-3. CSN5i-3 is a potent, selective and orally available inhibitor of CSN5.

-
GC62738
(R)-eIF4A3-IN-2
(R)-eIF4A3-IN-2 is a less active enantiomer of eIF4A3-IN-2.
-
GC70240
(R)-Fasiglifam
(R)-Fasiglifam is the isomer of Fasiglifam , and can be used as an experimental control.



