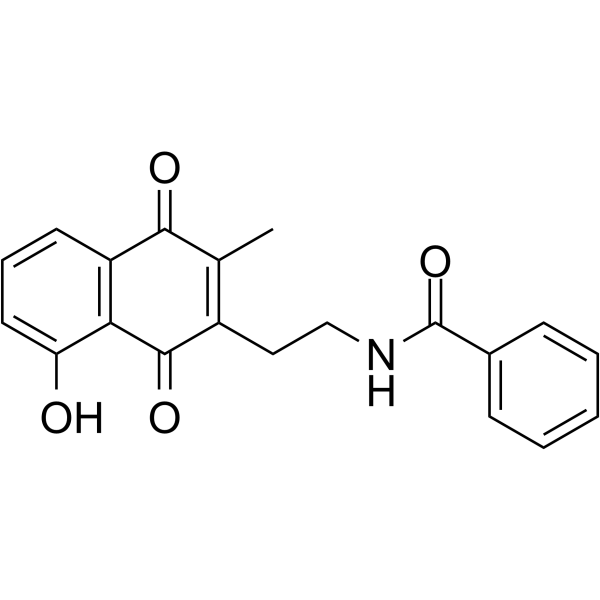
STAT
STAT (signal transducer and activator of transcription) are involved in the development and function of the immune system and play a role in maintaining immune tolerance and tumour surveillance.
Products for STAT
- Cat.No. Product Name Information
-
GC61807
(E/Z)-AG490
(E/Z)-AG490 ((E/Z)-Tyrphostin AG490) is a racemic compound of (E)-AG490 and (Z)-AG490 isomers. (E)-AG490 is a tyrosine kinase inhibitor that inhibits EGFR, Stat-3 and JAK2/3.

-
GC69836
(R,R)-VVD-118313
(R,R)-VVD-118313 is an isomer of VVD-118313. VVD-118313 is a selective JAK1 inhibitor that can block JAK1-dependent phosphorylation and cytokine signaling. VVD-118313 can be used for cancer research.

-
GC46503
2-(1,8-Naphthyridin-2-yl)phenol
2-NP
2-(1,8-Naphthyridin-2-yl)phenol is a selective enhancer of STAT1 transcription. 2-(1,8-Naphthyridin-2-yl)phenol can enhance the ability of IFN-γ to inhibit the proliferation of human breast cancer and fibrosarcoma cells.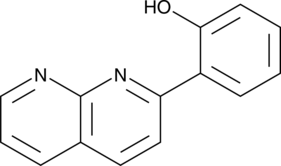
-
GC12138
5,15-DPP
5,15-Diphenylporphyrin,STAT3 Inhibitor VIII
5,15-DPP (5,15-DPP) is a selective STAT3-SH2 antagonist (IC50s of 0.28 μM and 10 μM for STAT3 and STAT1, respectively).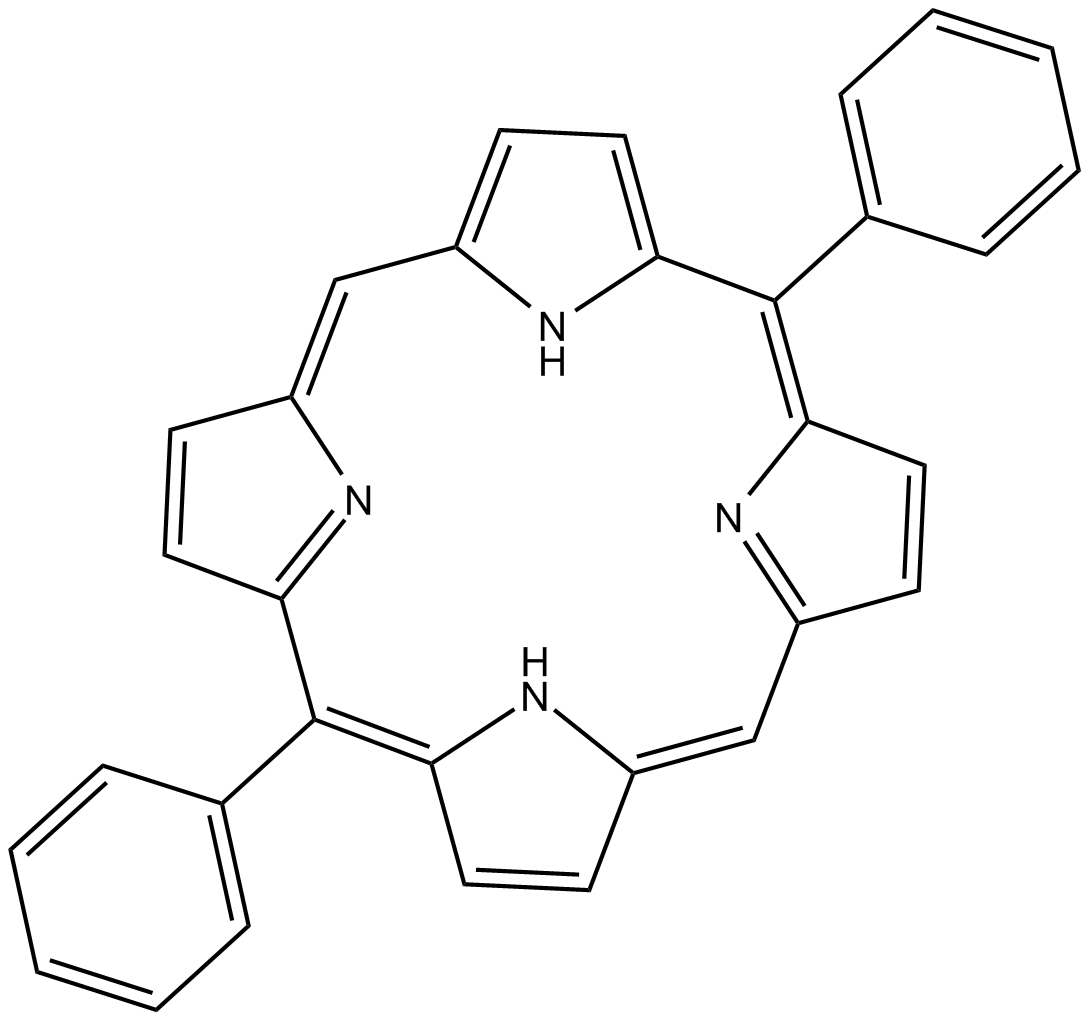
-
GC62272
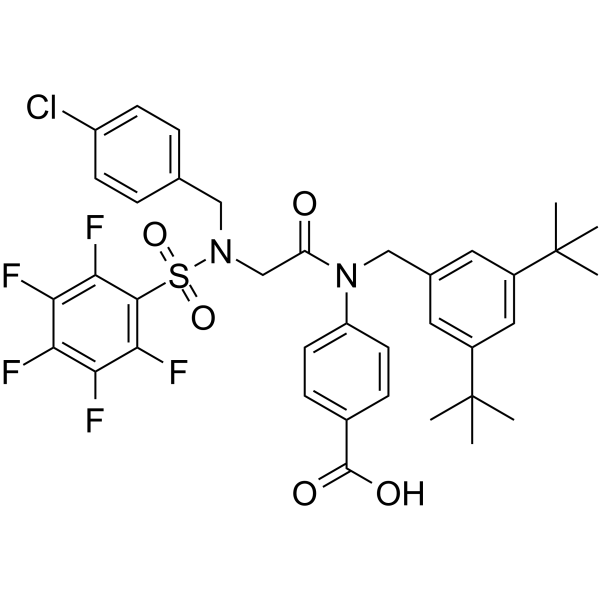
AC-4-130
AC-4-130 is a potent STAT5 SH2 domain inhibitor. AC-4-130 directly binds to STAT5 and disrupts STAT5 activation, dimerization, nuclear translocation, and STAT5-dependent gene transcription. AC-4-130 induces cell cycle arrest and apoptosis in FLT3-ITD-driven leukemic cells. AC-4-130 has anti-cancer activity and can efficiently block pathological levels of STAT5 activity in acute myeloid leukemia (AML).
-
GC13854
AG-490 (Tyrphostin B42)
Tyrphostin AG-490
AG-490 (Tyrphostin B42) (Tyrphostin AG-490 (Tyrphostin B42)) is a tyrosine kinase inhibitor that inhibits EGFR, Stat-3 and JAK2/3.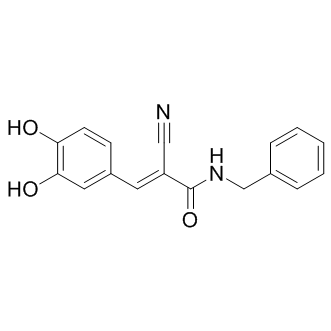
-
GN10705
Alantolactone
(+)-Alantolactone, NSC 333843, NSC 93131
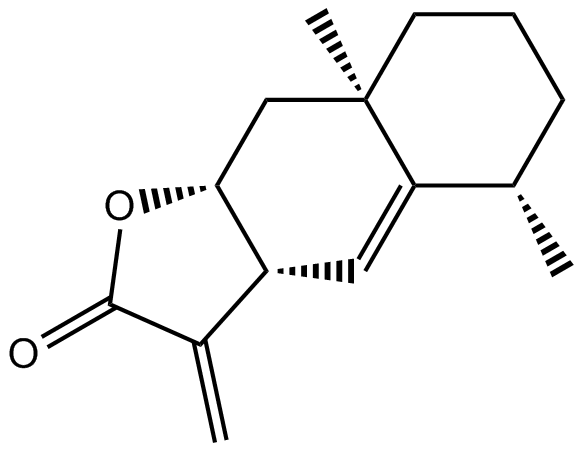
-
GC60585
Angoline
Angoline is a potent and selective IL6/STAT3 signaling pathway inhibitor with an IC50 of 11.56 μM. Angoline inhibits STAT3 phosphorylation and its target gene expression, and inhibits cancer cell proliferation.
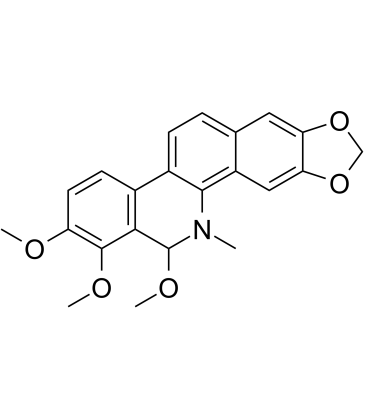
-
GC60586
Angoline hydrochloride
Angoline hydrochloride is a potent and selective IL6/STAT3 signaling pathway inhibitor with an IC50 of 11.56 μM. Angoline hydrochloride inhibits STAT3 phosphorylation and its target gene expression, and inhibits cancer cell proliferation.
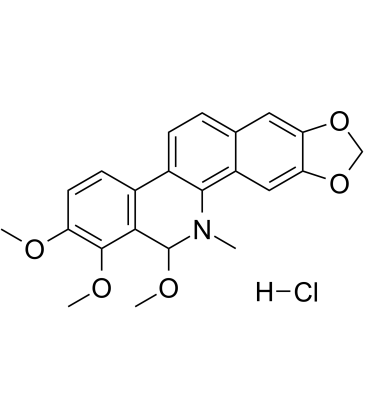
-
GC10439
APTSTAT3-9R
STAT3 inhibitor

-
GC35395
Arnicolide D
ARD
Arnicolide D is a sesquiterpene lactone isolated from Centipeda minima. Arnicolide D modulates the cell cycle, activates the caspase signaling pathway and inhibits the PI3K/AKT/mTOR and STAT3 signaling pathways. Arnicolide D inhibits Nasopharyngeal carcinoma (NPC) cell viability in a concentration- and time-dependent manner.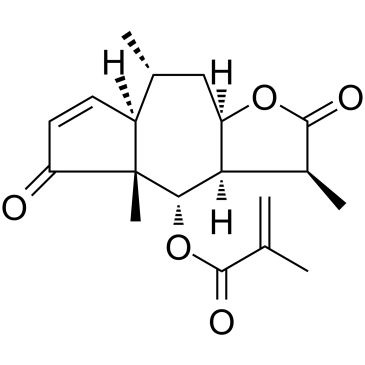
-
GC10889
Artesunate
Artesunic Acid, NSC 712571, WR 256283
Derivative of the natural product artemisinin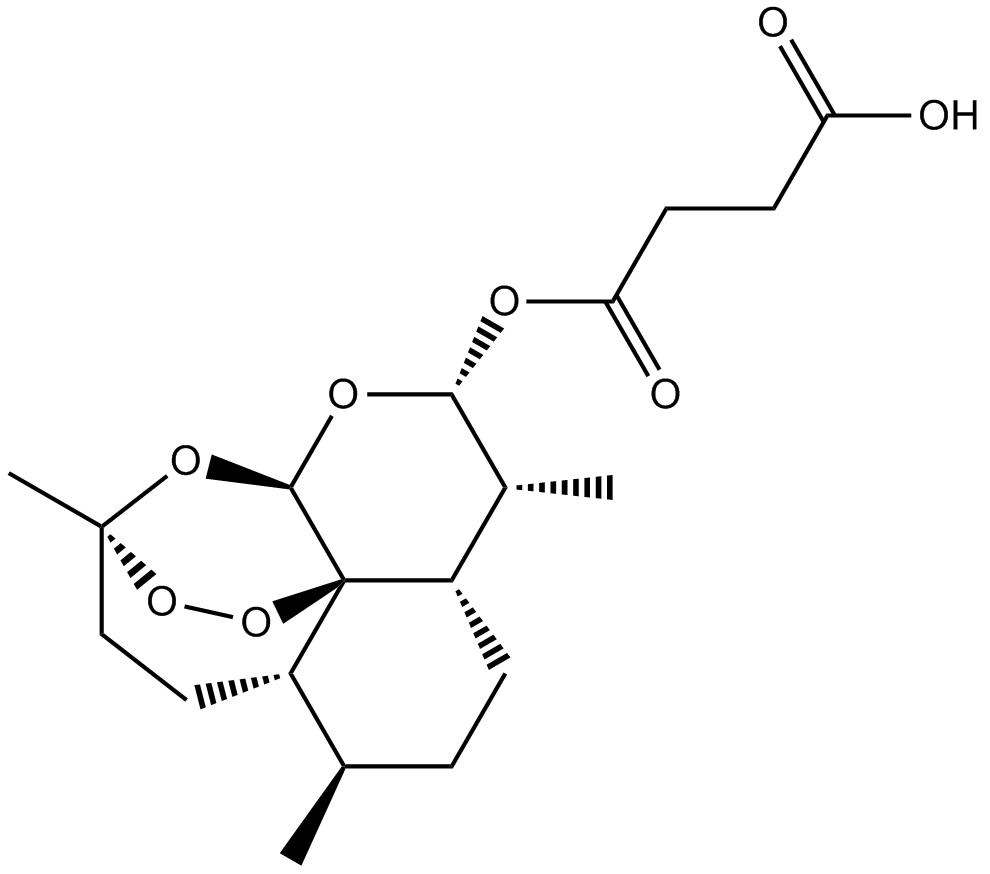
-
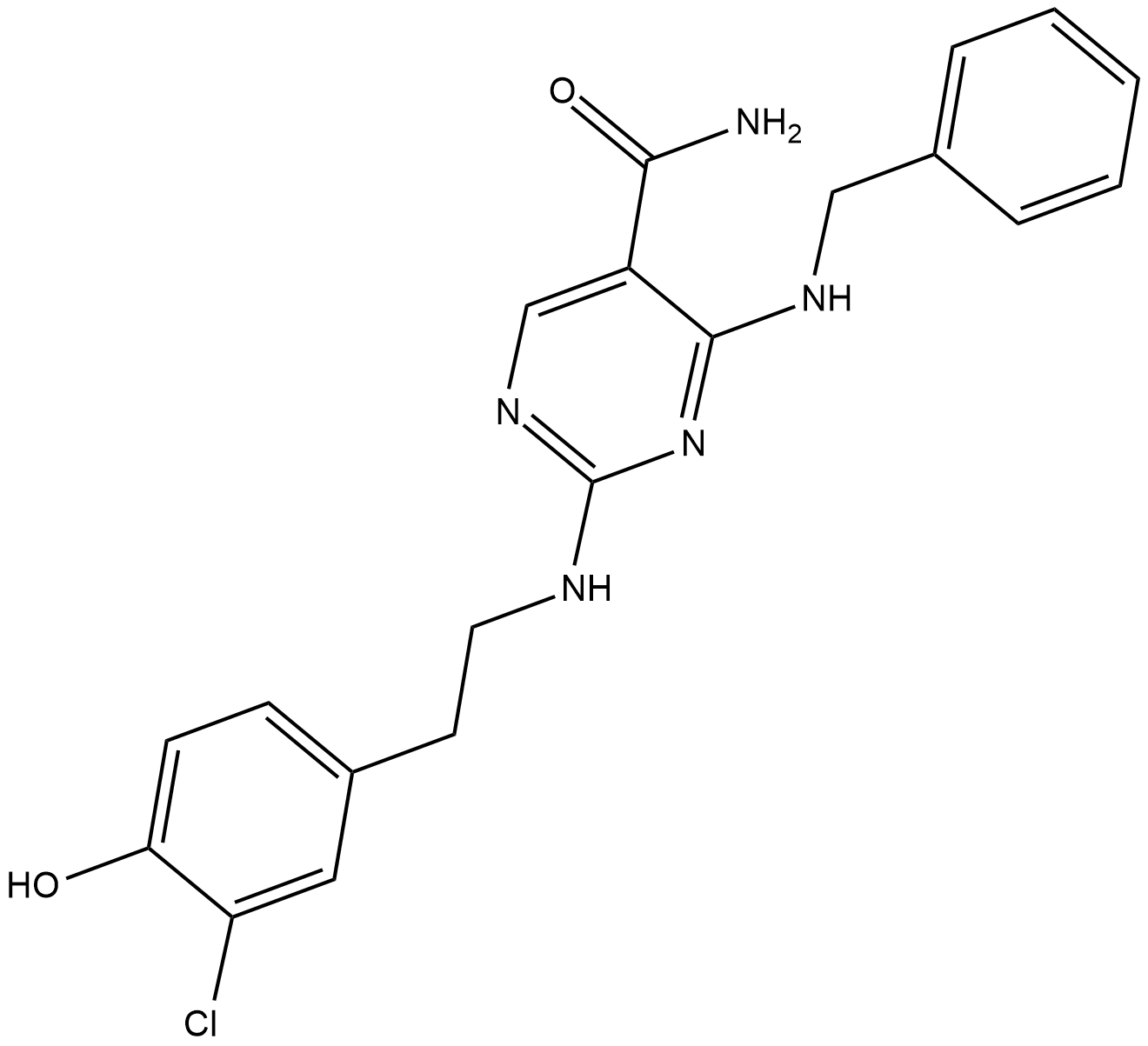
GC19039
AS1517499
A potent STAT6 inhibitor
-
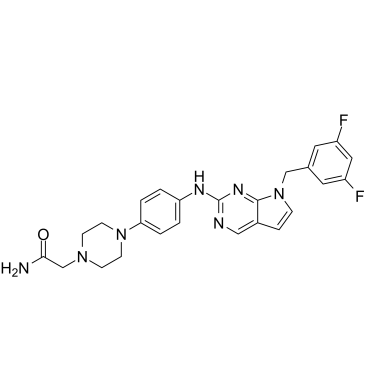
GC61574
AS1810722
AS1810722 is an orally active and potent STAT6 inhibitor with an IC50 of 1.9 nM.
-
GC38736
AS2863619
A dual inhibitor of Cdk8 and Cdk19
-
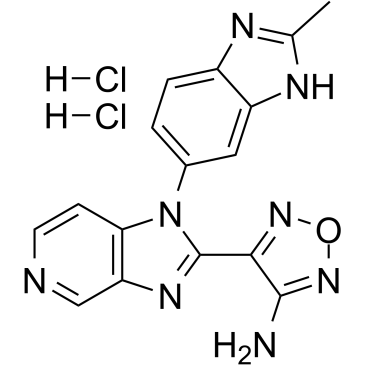
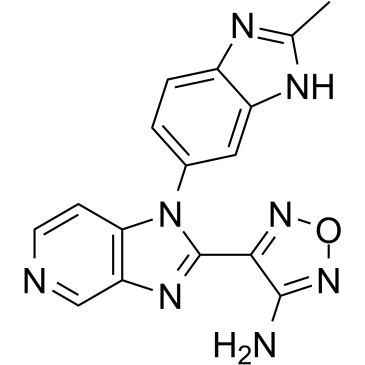
GC38737
AS2863619 free base
AS2863619 free base enables conversion of antigen-specific effector/memory T cells into Foxp3+ regulatory T (Treg) cells for the treatment of various immunological diseases.
-
GC40715
Ascochlorin
Antibiotic LL-Z1272γ, Ilicicolin D, NSC 287492
Ascochlorin is an isoprenoid antibiotic and antiviral that has diverse effects on mammalian cells.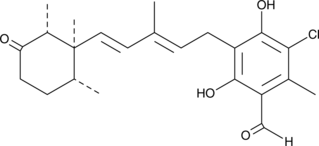
-
GN10627
Atractylenolide I
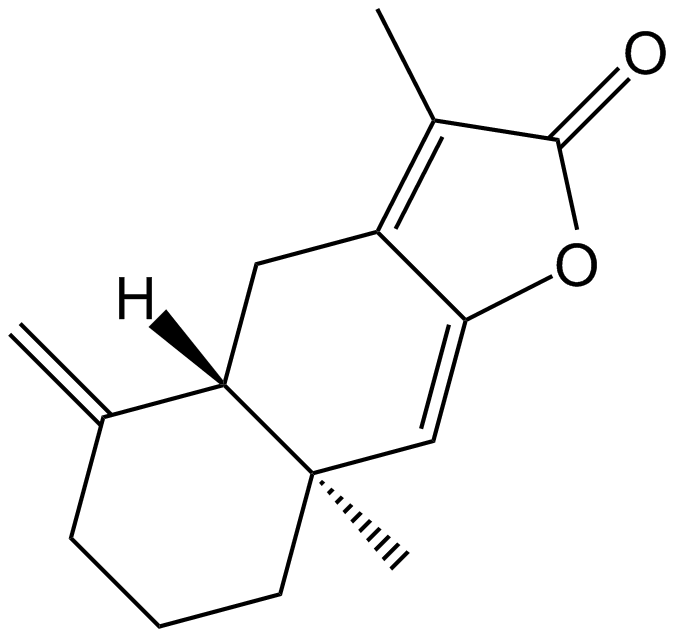
-
GC18126
Balsalazide
anti-inflammatory drug
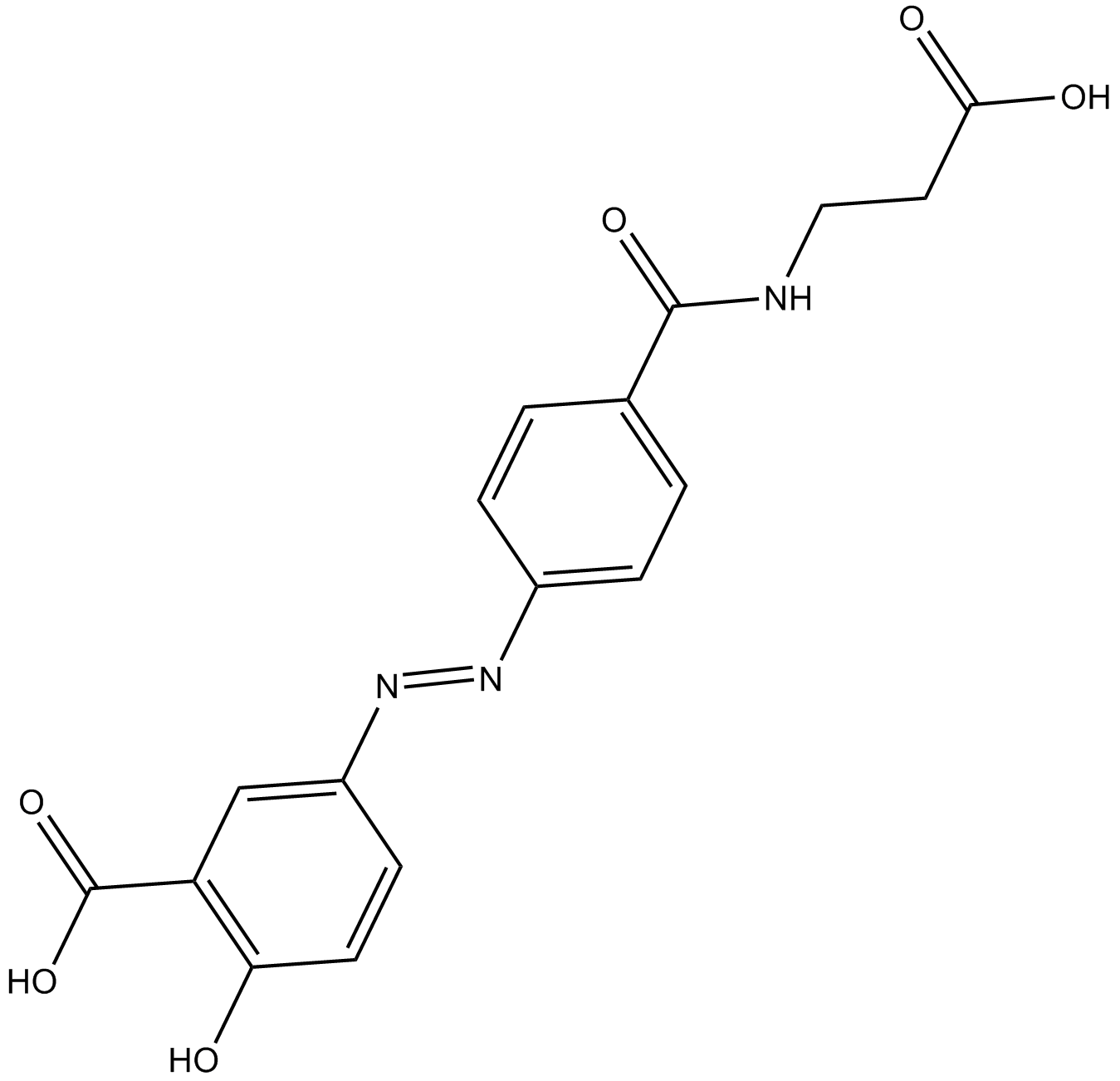
-
GC35466
Balsalazide sodium hydrate
Balsalazide sodium hydrate could suppress colitis-associated carcinogenesis through modulation of IL-6/STAT3 pathway.
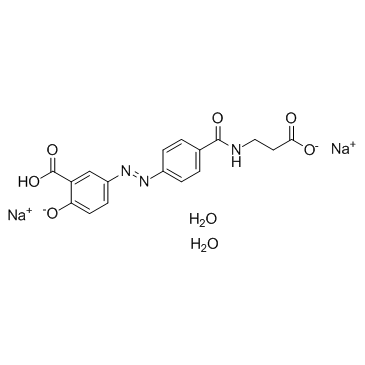
-
GC34063
BP-1-102
A STAT3 inhibitor
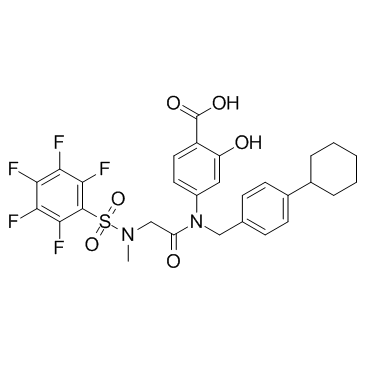
-
GC42974
Brassinin
BSN
Brassinin (BSN) is a phytoalexin isolated from B.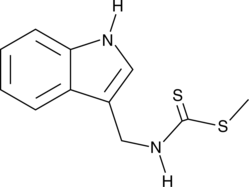
-
GC35554
Brevilin A
6-O-Angeloylprenolin, Brevelin A
A sesquiterpene lactone with anticancer activity
-
GC34076
C188-9
TTI-101
A STAT3 inhibitor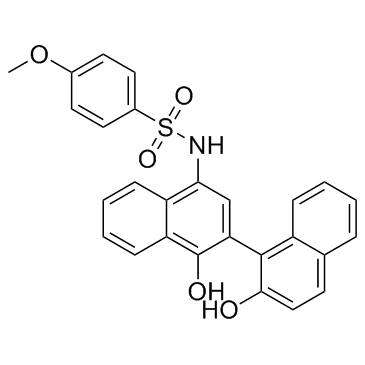
-
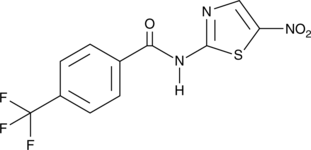
GC47061
CAY10763
A dual inhibitor of IDO1 and STAT3 activation
-
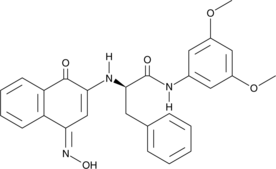
GC49139
CAY10784
A STAT3 inhibitor
-
GC35651
Cenisertib
AS-703569; R-763
Cenisertib (AS-703569) is an ATP-competitive multi-kinase inhibitor that blocks the activity of Aurora-kinase-A/B, ABL1, AKT, STAT5 and FLT3. Cenisertib induces major growth-inhibitory effects by blocking the activity of several different molecular targets in neoplastic mast cells (MC). Cenisertib inhibits tumor growth in xenograft models of pancreatic, breast, colon, ovarian, and lung tumors and leukemia.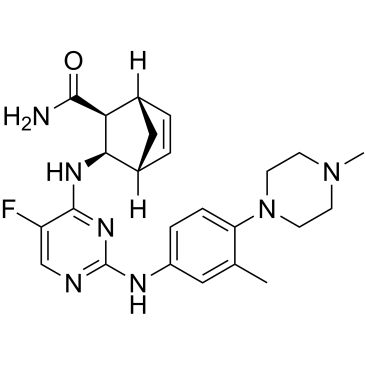
-
GC34238
CMD178
CMD178 is a lead peptide that consistently reduced the expression of Foxp3 and STAT5 induced by IL-2/s IL-2Rα signaling and inhibits Treg cell development.
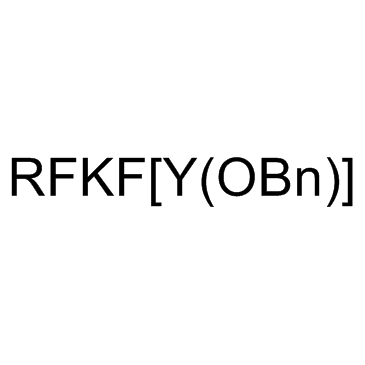
-
GC35715
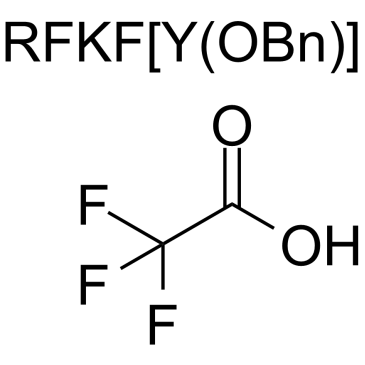
CMD178 TFA
CMD178 TFA is a lead peptide that consistently reduced the expression of Foxp3 and STAT5 induced by IL-2/s IL-2Rα signaling and inhibits Treg cell development.
-
GC14854
Colivelin
Colivelin (CLN) is A brain-permeable neuroprotective peptide that has effective long-term effects on Aβ deposition, neuronal apoptosis and synaptic plasticity defects in neurodegenerative diseases.

-
GC35720
Colivelin TFA
Colivelin TFA is a brain penetrant neuroprotective peptide and a potent activator of STAT3, suppresses neuronal death by activating STAT3?in vitro.

-
GC13279
Corylifol A
Corylinin
STAT3 inhibitor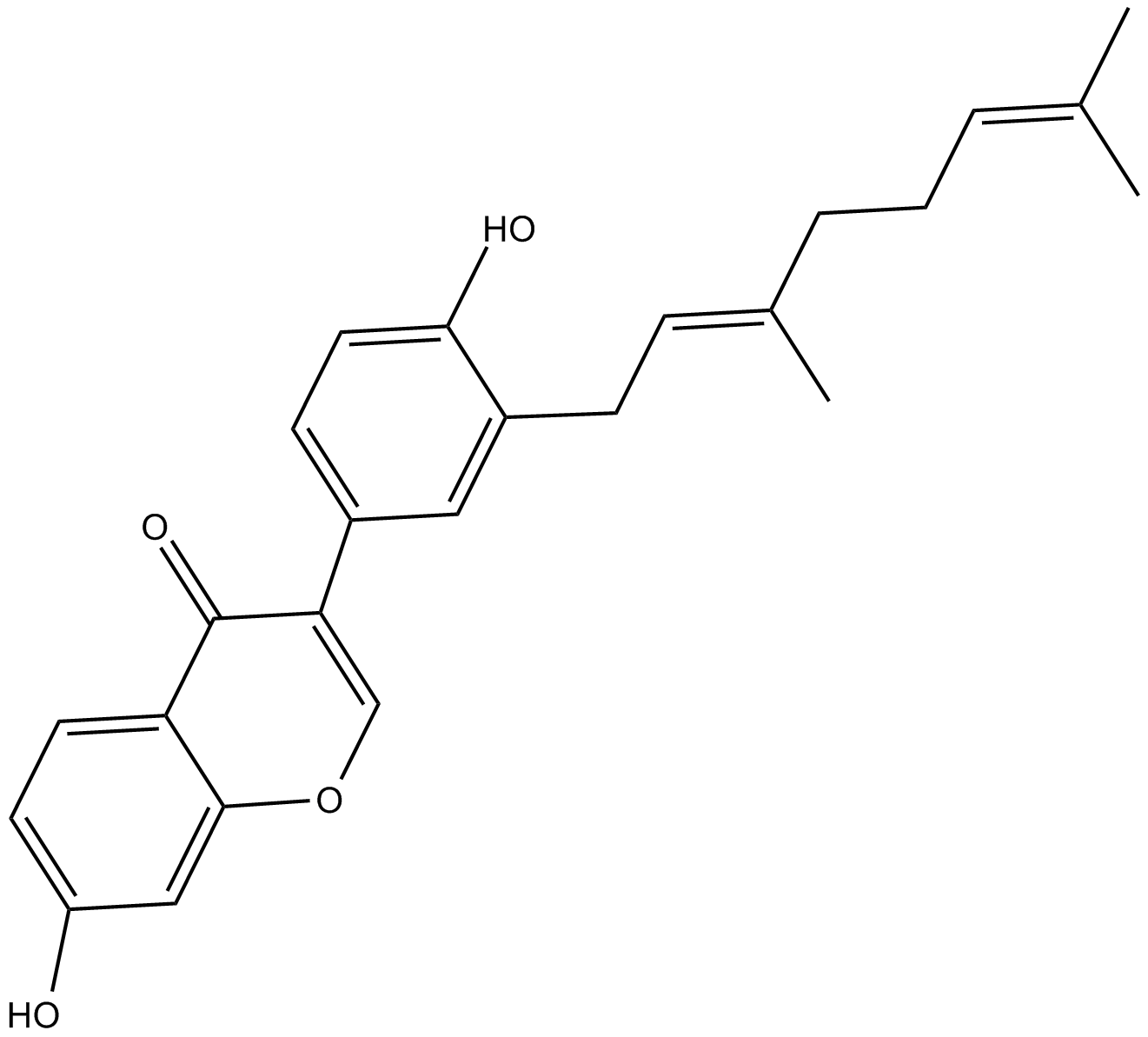
-
GN10501
Cryptotanshinone
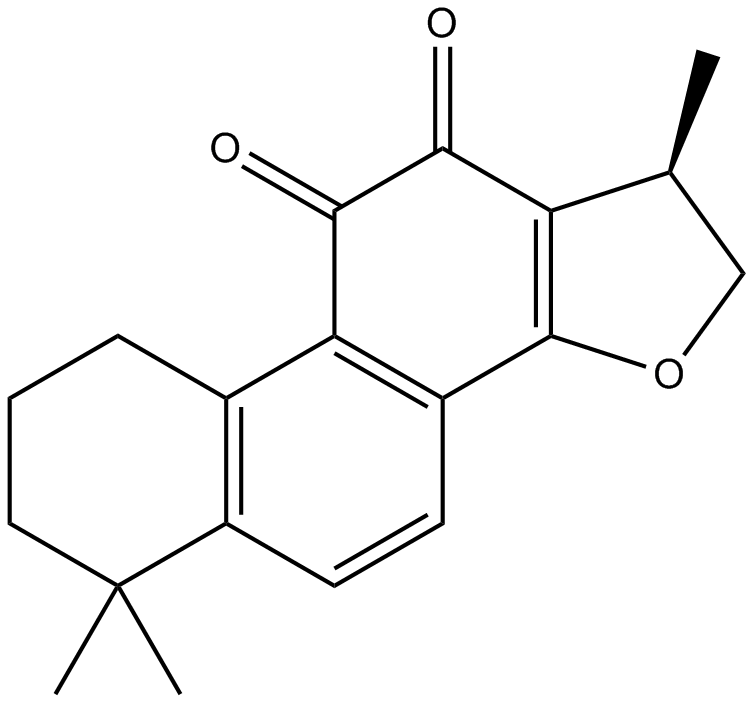
-
GC13347
Cucurbitacin I
Elatericin B; JSI-124; NSC-521777
An inhibitor STAT3/JAK signaling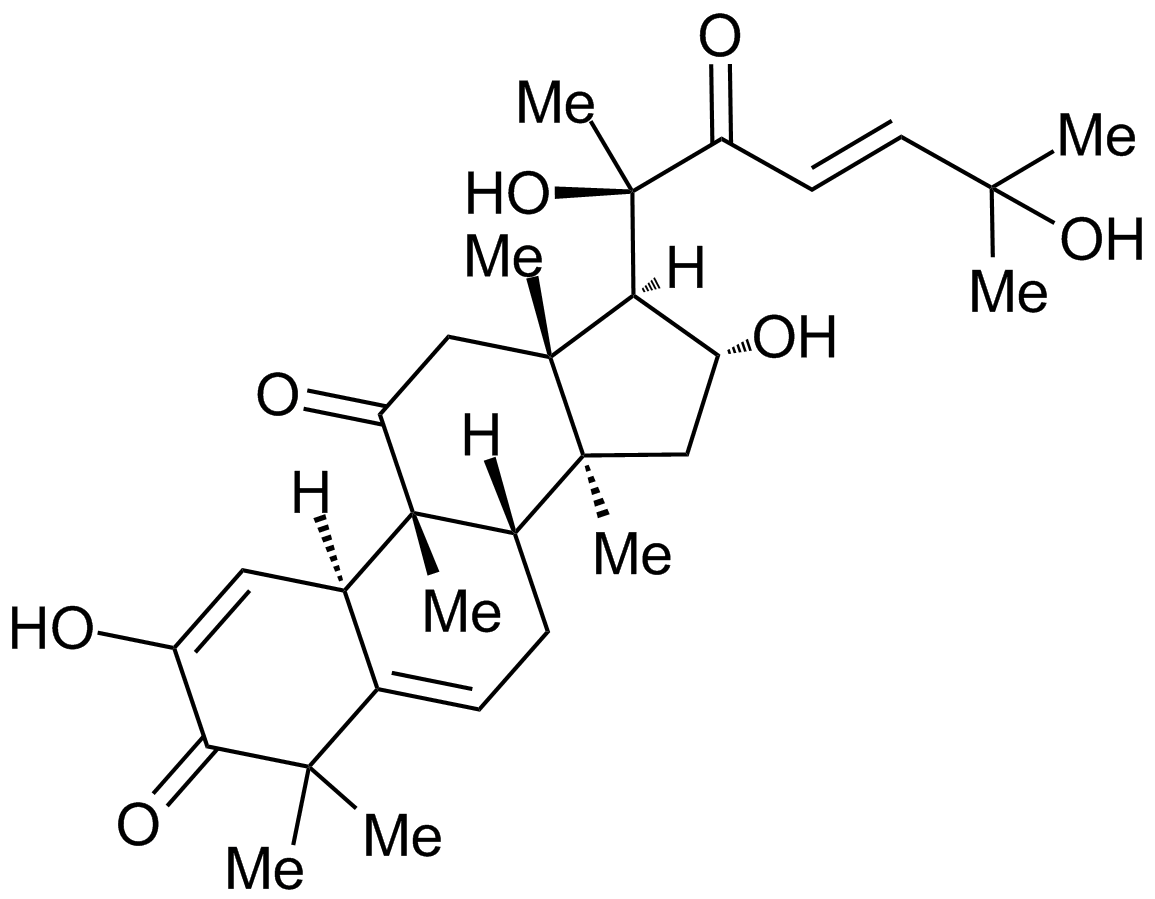
-
GN10442
Curculigoside
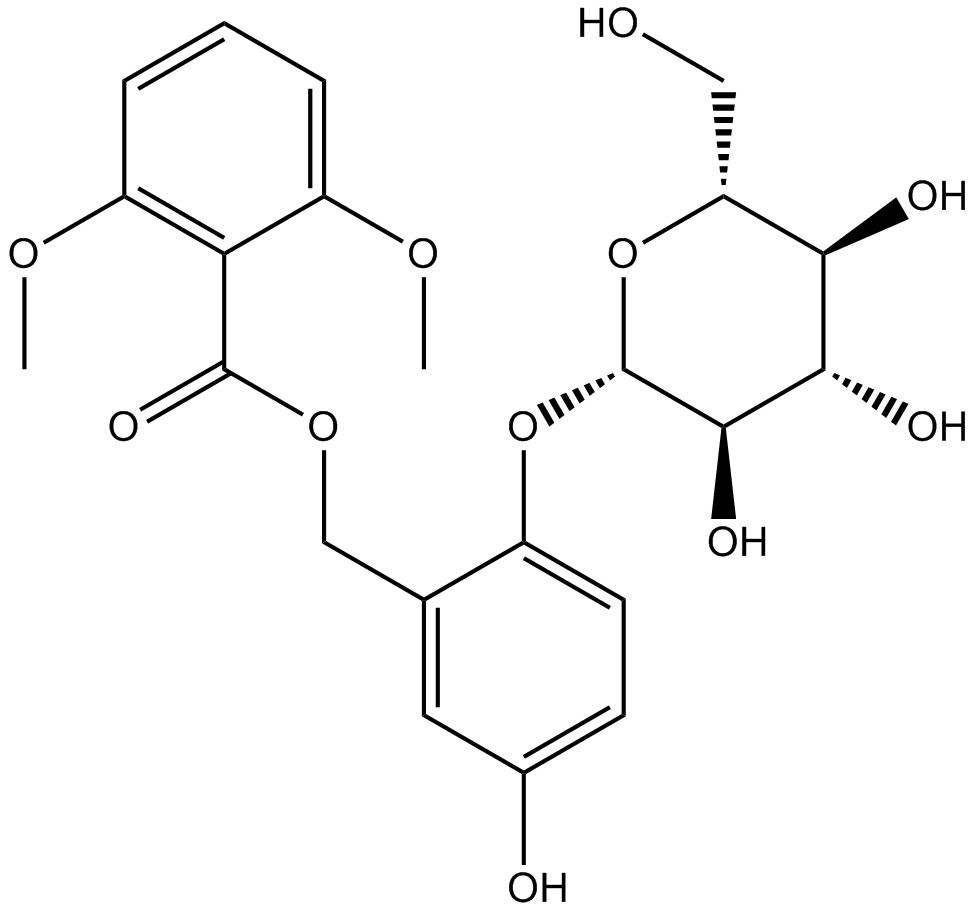
-
GC65020
Danvatirsen
AZD9150
Danvatirsen is an antisense oligonucleotide targeting STAT3 with potential antitumor activity. Danvatirsen binds to STAT3 mRNA, thereby inhibiting translation of the transcript. Suppression of STAT3 expression induces tumor cell apoptosis and decreases tumor cell growth.
-
GC43406
Delphinidin (chloride)
Ephdine
Delphinidin (chloride) is an anthocyanidin, a natural plant pigment which serves as the precursor of certain anthocyanins that provide the blue-red colors of flowers, fruits, and red wine.
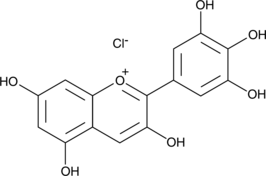
-
GC34101
Dihydroisotanshinone I
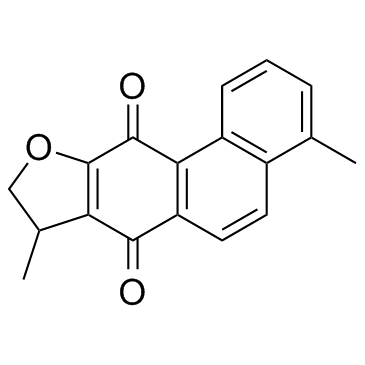
-
GN10115
Diosgenin
Nitogenin, NSC 33396, 3β-hydroxy-5-Spirostene
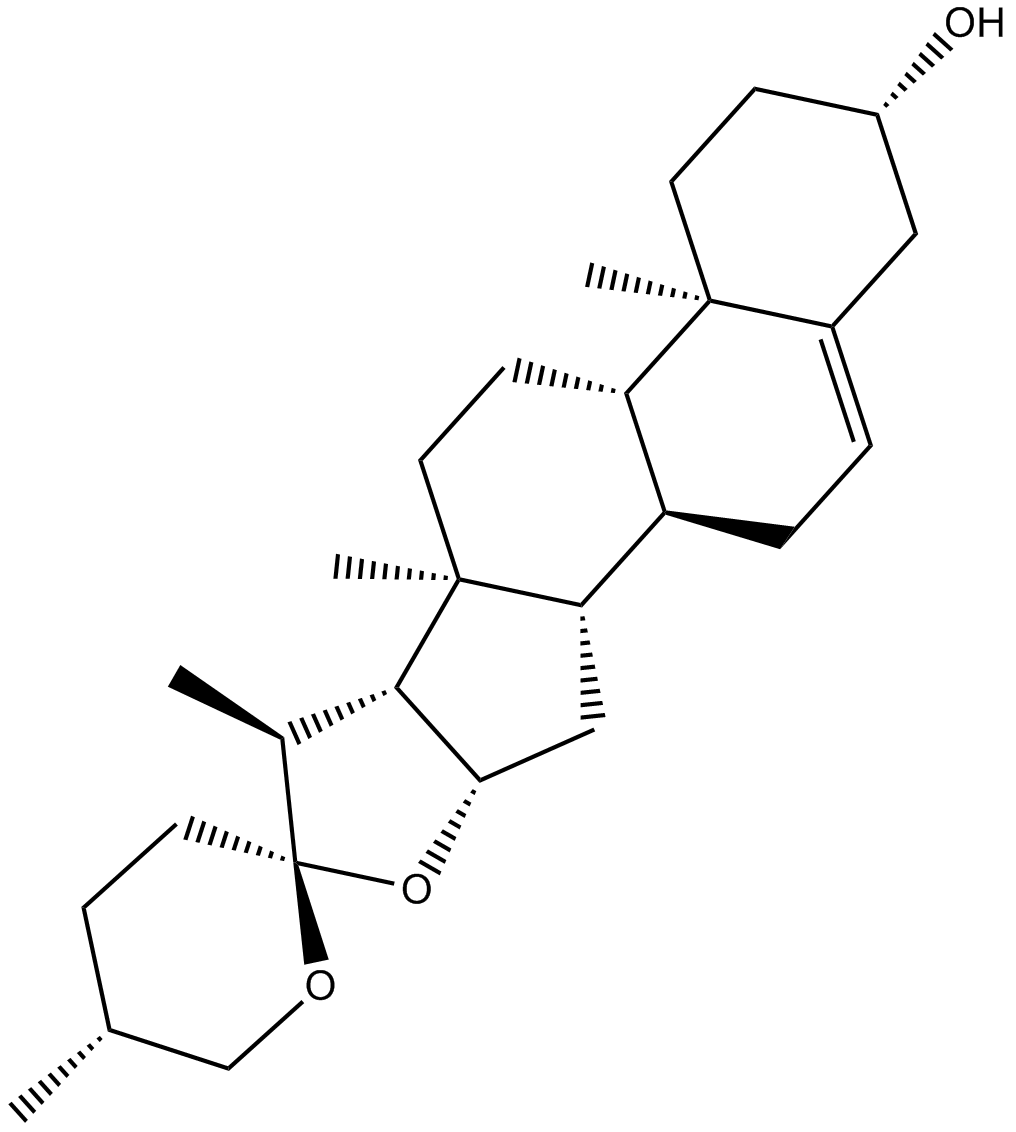
-
GC33170
ENMD-119 (ENMD 1198)
IRC-110160
ENMD-119 (ENMD 1198) (IRC-110160), an orally active microtubule destabilizing agent, is a 2-methoxyestradiol analogue with antiproliferative and antiangiogenic activity. ENMD-119 (ENMD 1198) is suitable for inhibiting HIF-1alpha and STAT3 in human HCC cells and leads to reduced tumor growth and vascularization.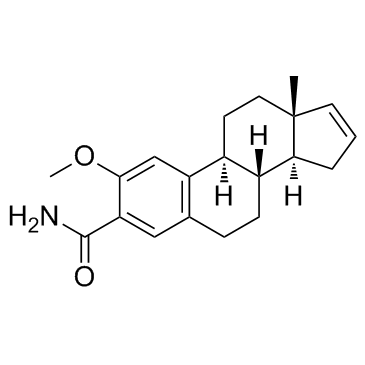
-
GC36016
Eupalinolide K
Eupalinolide K, a sesquiterpene lactones compound from Eupatorium lindleyanum, is a STAT3 inhibitor. Eupalinolide K is a Michael reaction acceptor (MRA) .
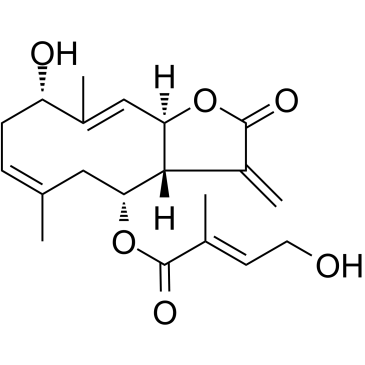
-
GC16875
FLLL32
JAK2 Inhibitor X
STAT3 inhibitor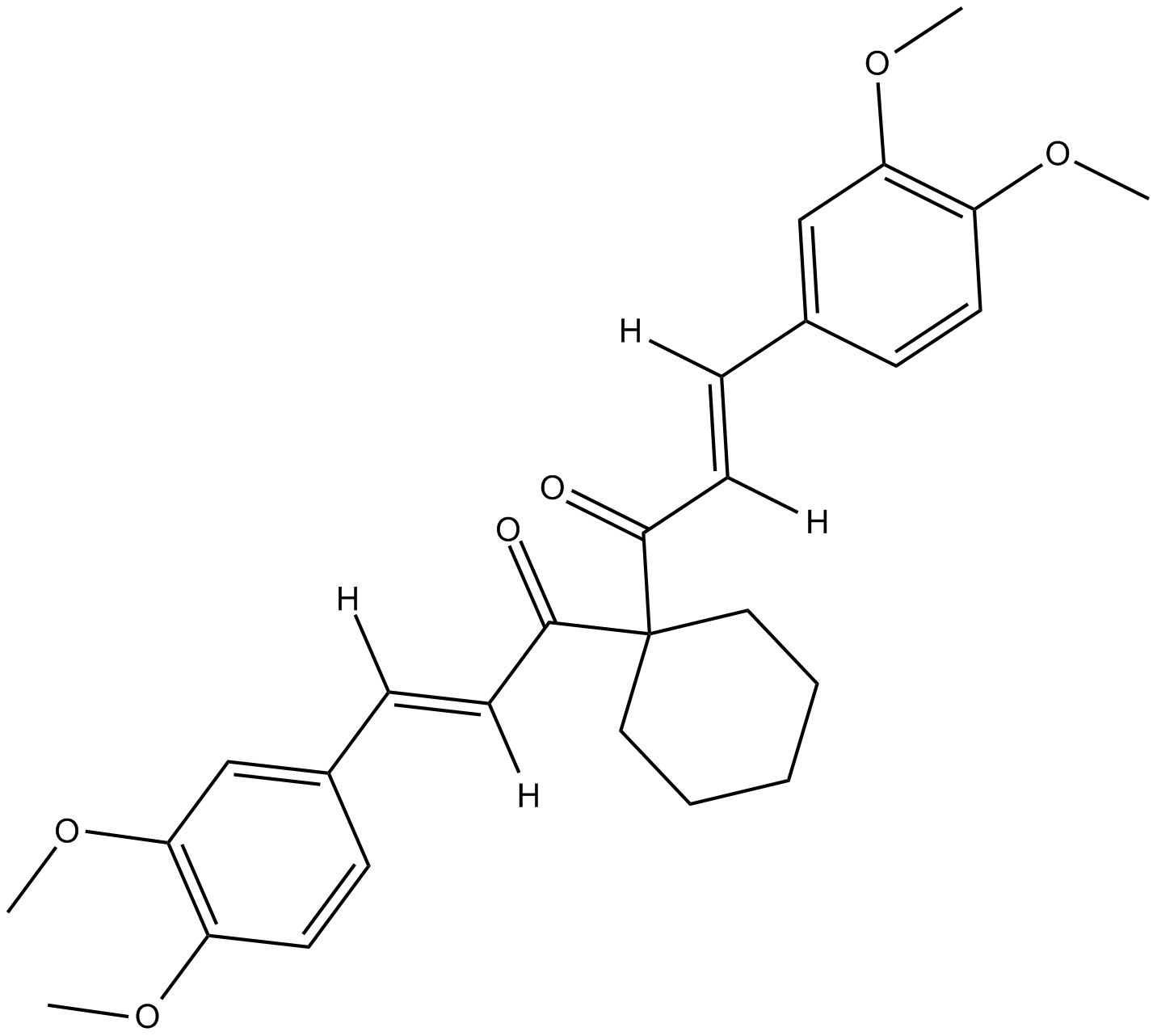
-
GC14144
Fludarabine
2-fluoro-ara-A, NSC 118218
DNA synthsis inhibitor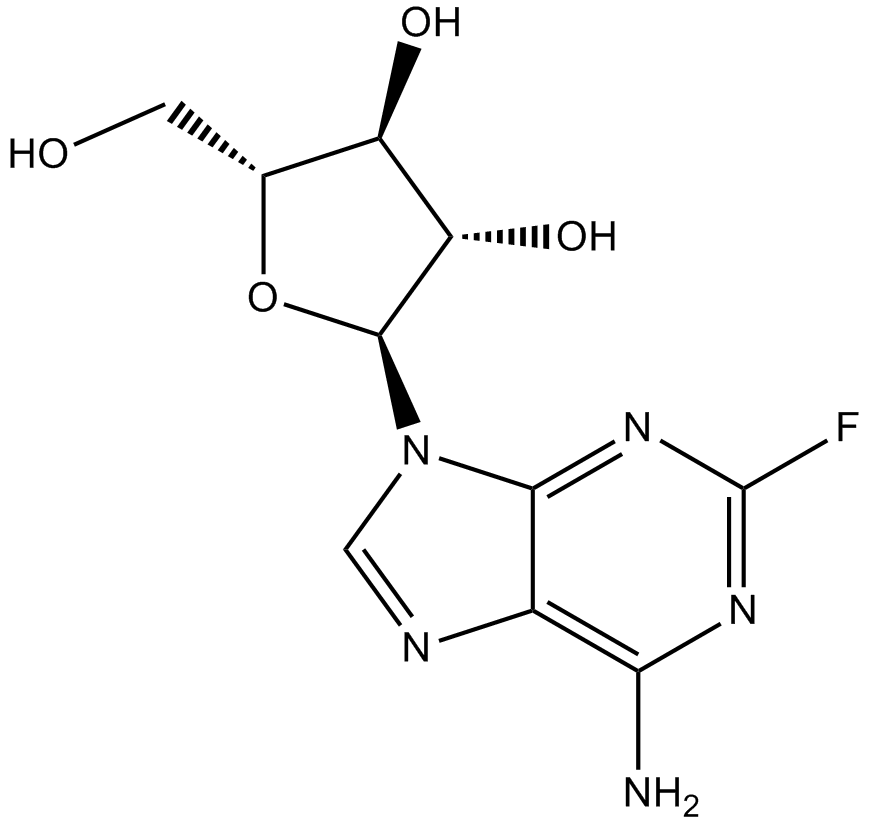
-
GC15134
Fludarabine Phosphate (Fludara)
Fludura
Fludarabine (phosphate) is an analogue of adenosine and deoxyadenosine, which is able to compete with dATP for incorporation into DNA and inhibit DNA synthesis.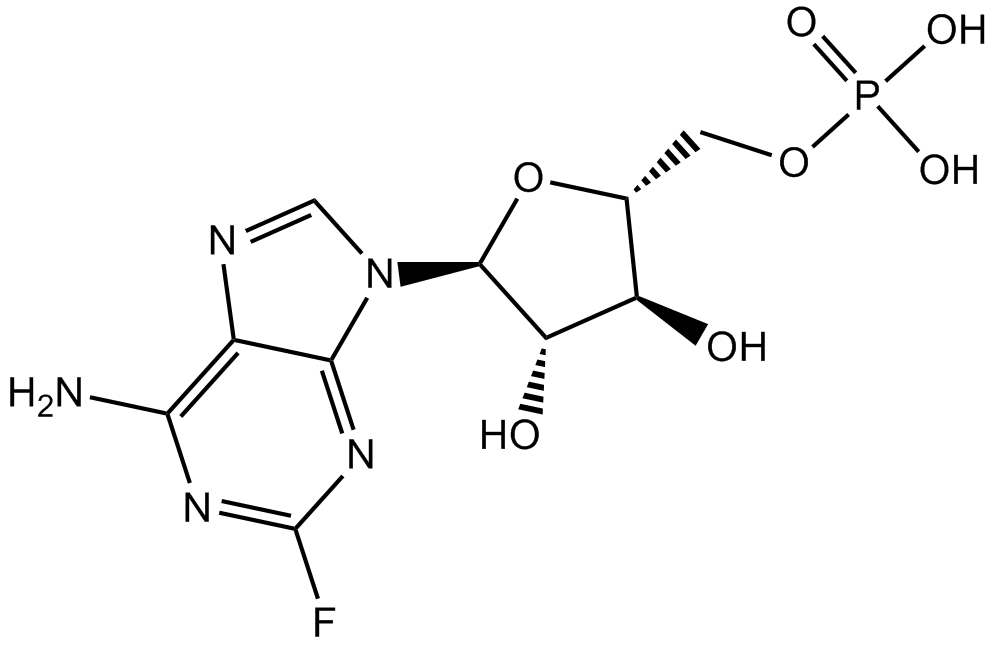
-
GC38044
Fraxinellone
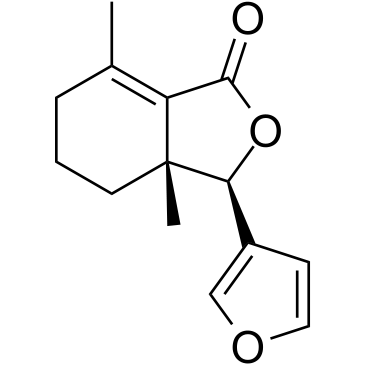
-
GC13285
Galiellalactone
inhibits IL-6-mediated JAK/STAT signal transduction
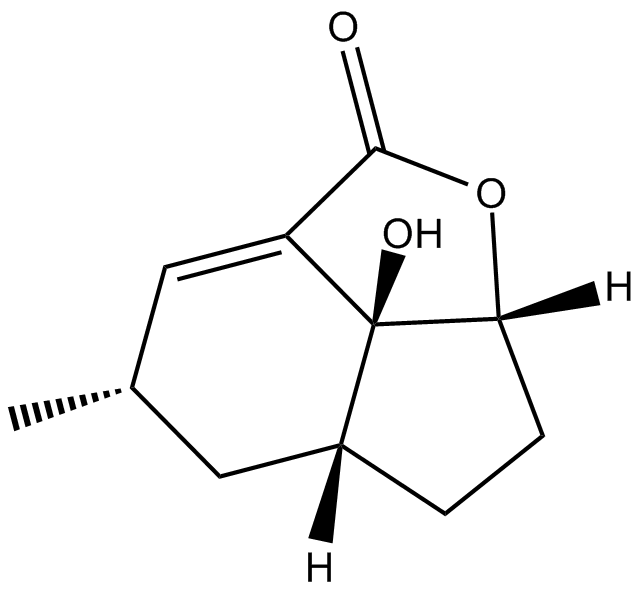
-
GN10696
Garcinone C
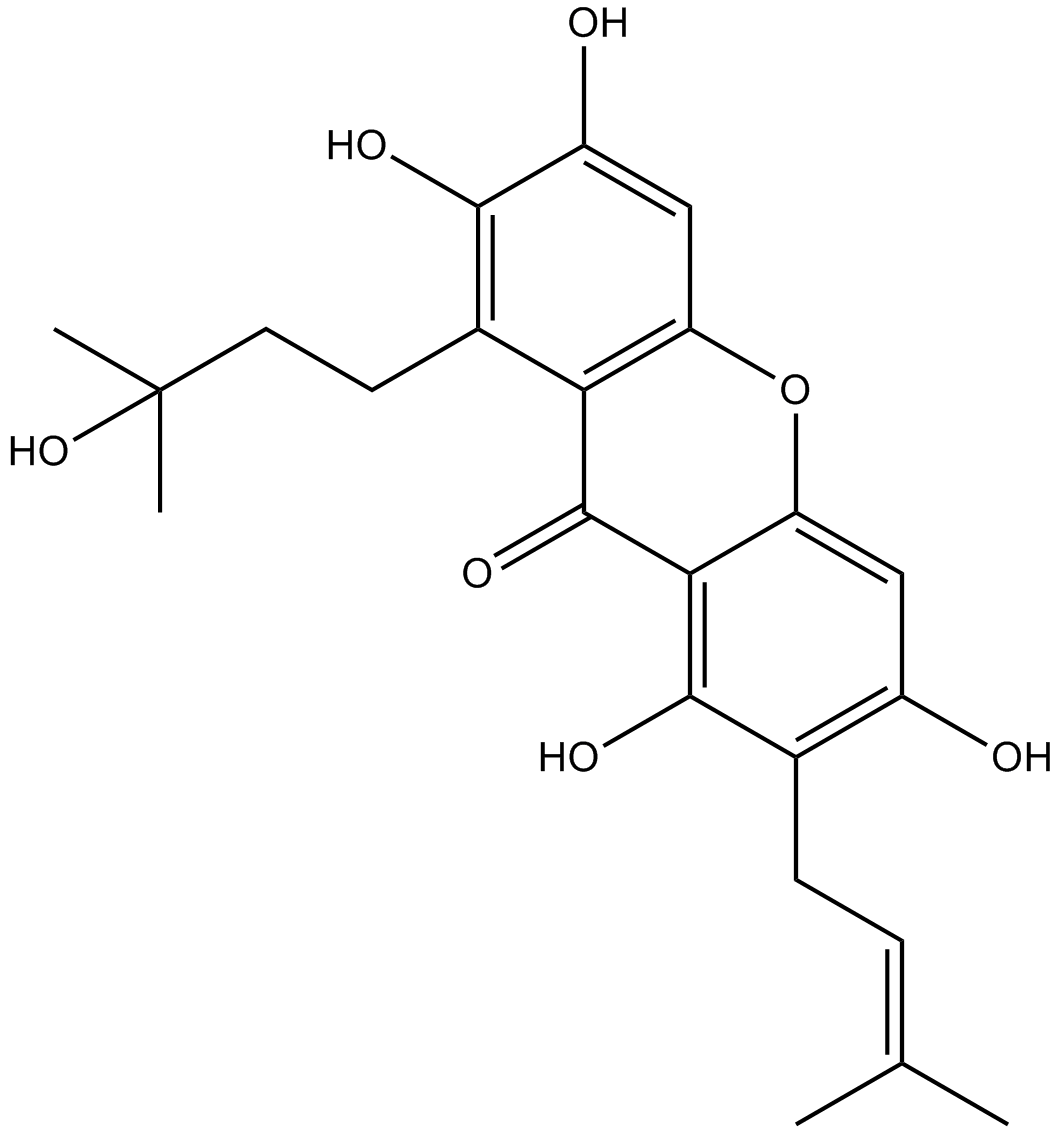
-
GN10741
Garcinone D
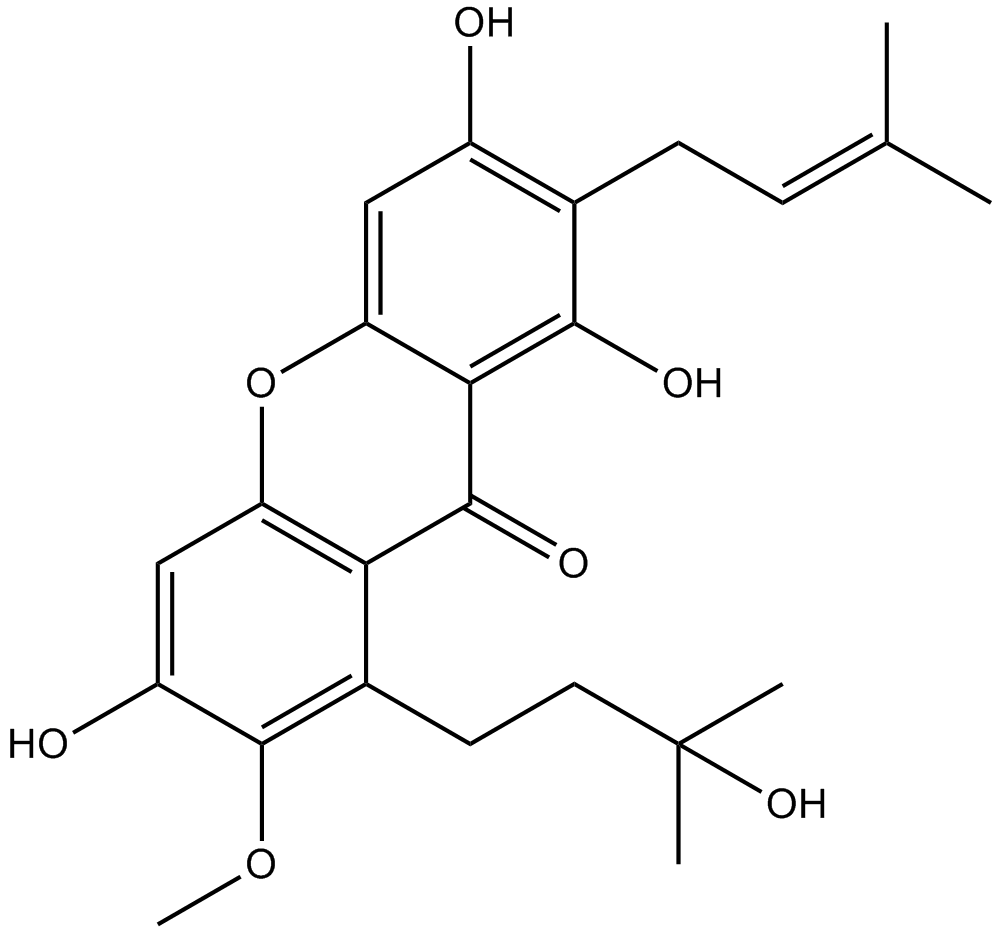
-
GC36174
Golotimod
SCV 07; Gamma-D-glutamyl-L-tryptophan
Golotimod (SCV-07), an immunomodulating peptide with antimicrobial activity, significantly increases the efficacy of antituberculosis therapy, stimulates thymic and splenic cell proliferation, and improves macrophage function.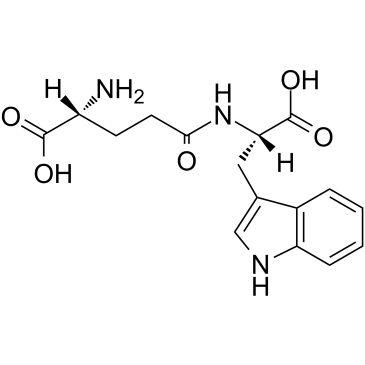
-
GC36175
Golotimod (TFA)
SCV 07 TFA; Gamma-D-glutamyl-L-tryptophan TFA
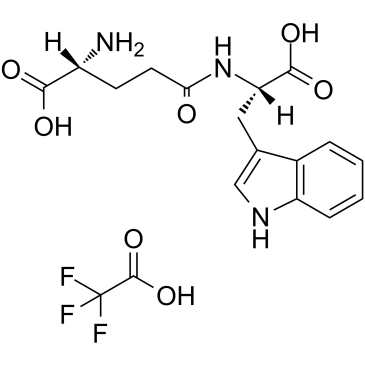
-
GC38386
Golotimod hydrochloride
SCV 07 hydrochloride; Gamma-D-glutamyl-L-tryptophan hydrochloride
Golotimod hydrochloride (SCV 07 hydrochloride), an immunomodulating peptide with antimicrobial activity, significantly increases the efficacy of antituberculosis therapy, stimulates thymic and splenic cell proliferation, and improves macrophage function.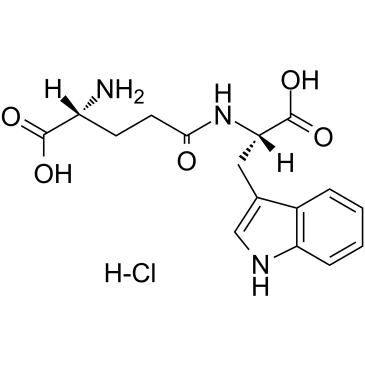
-
GC50343
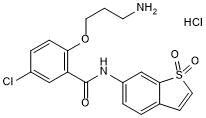
HJC 0416 hydrochloride
STAT3 inhibitor
-
GC32807
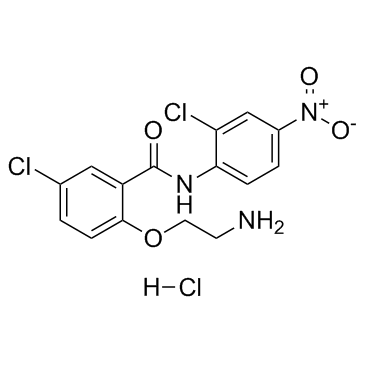
HJC0152 hydrochloride
An orally bioavailable inhibitor of STAT3
-
GC16208
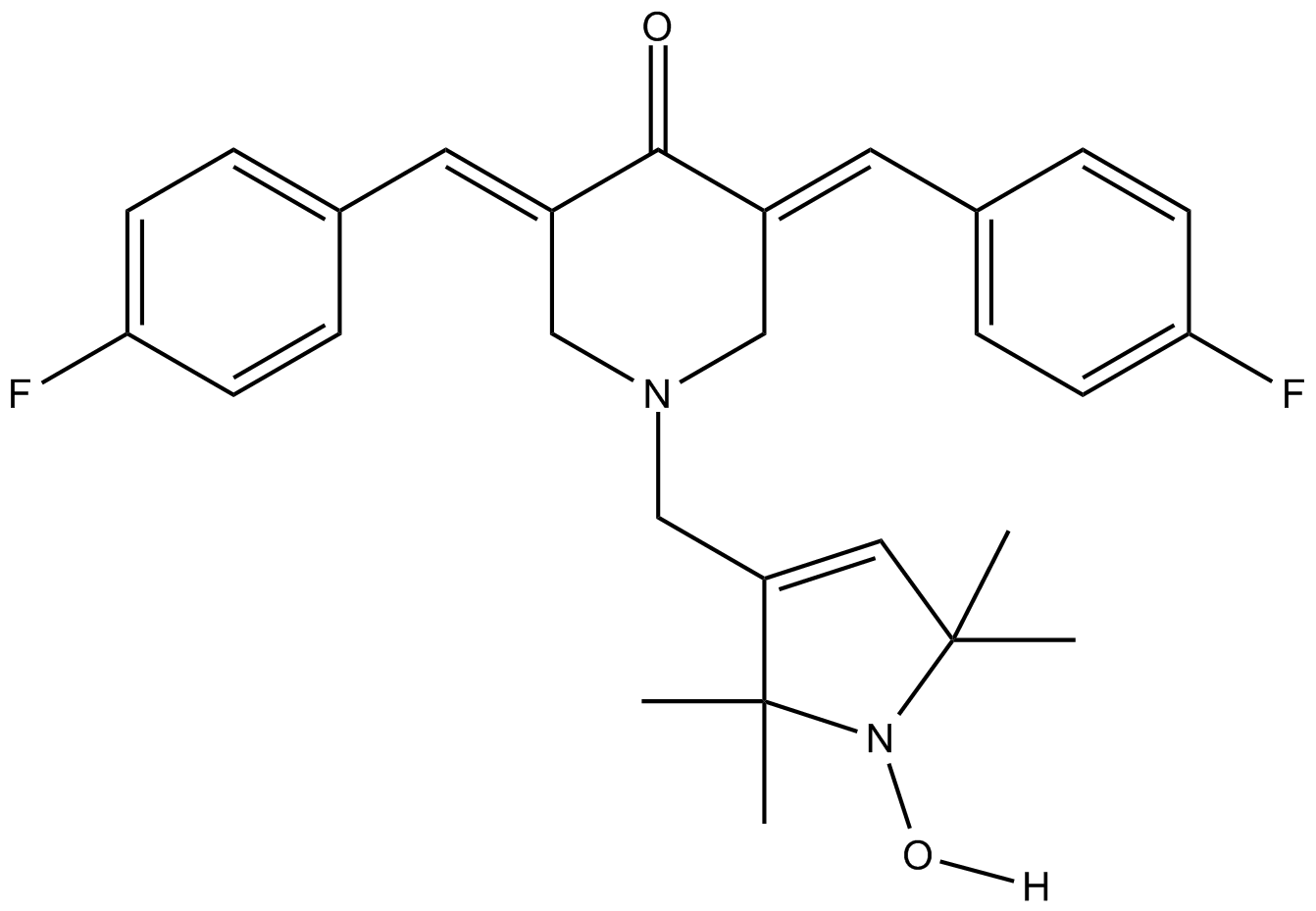
HO-3867
STAT3 inhibitor, selective
-
GN10742
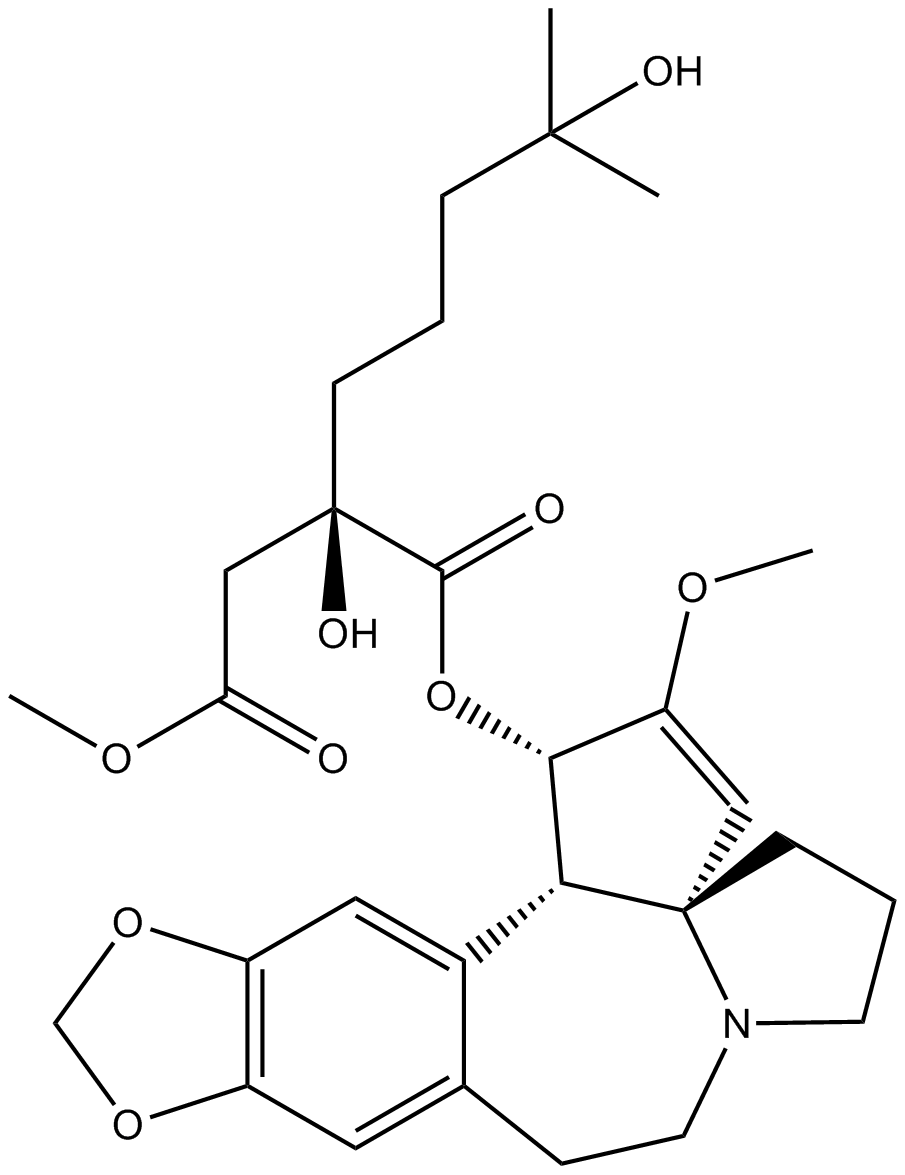
Homoharringtonine
NSC 141633, Omacetaxine Mepesuccinate
-
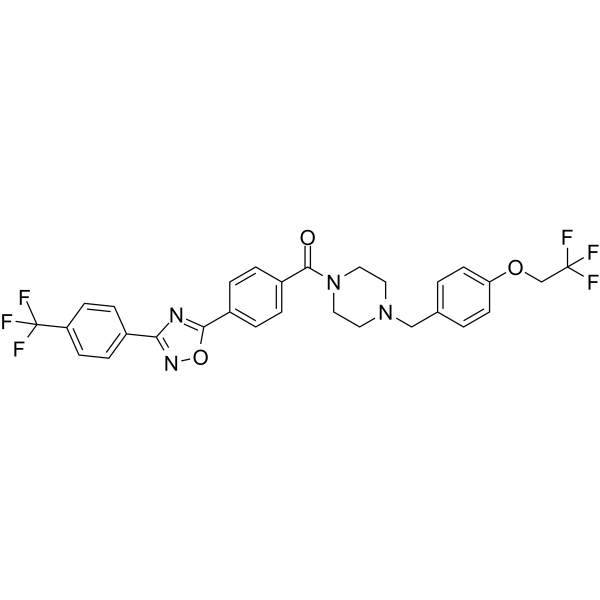
GC67958
HP590
-
GC32862
inS3-54A18
inS3-54A18 is a potent STAT3 inhibitor, with anti-cancer properties.

-
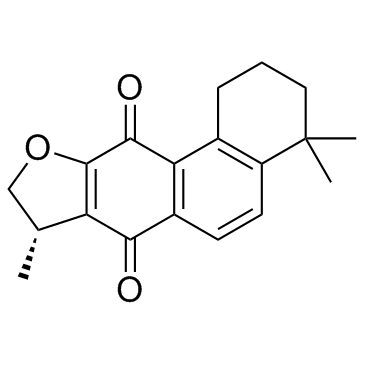
GC34178
Isocryptotanshinone
-
GC69300
IST5-002
N6-Benzyladenosine-5'-phosphate
IST5-002 is an effective inhibitor of Stat5a/b, selectively inhibiting the transcriptional activity of Stat5a/b with IC50 values of 1.5 μM (Stat5a) and 3.5 μM (Stat5b). IST5-002 induces apoptosis and death in prostate cancer cells and chronic myeloid leukemia (CML) cells. IST5-002 can be used for research on prostate cancer and CML.

-
GC15731
L002
p300/CBP Inhibitor VI, NSC 764414
p300 inhibitor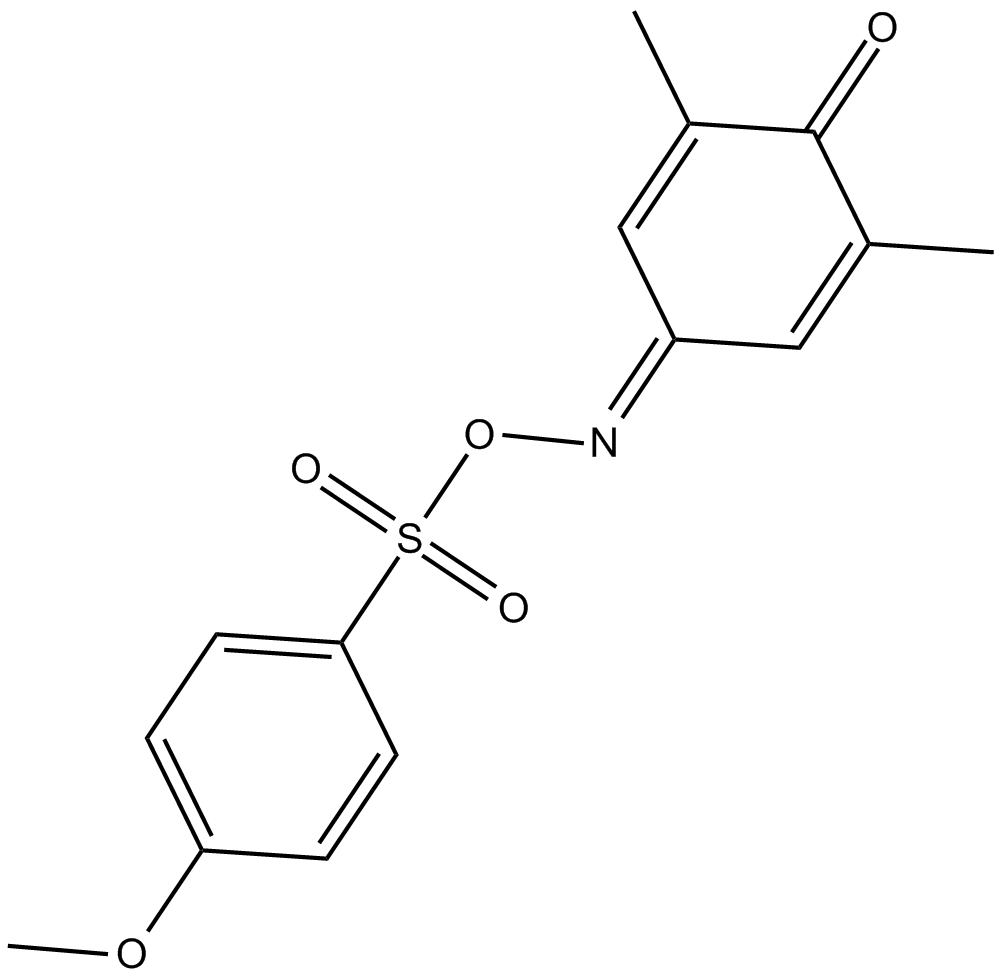
-
GC69384
LLL12
LLL12 is a small molecule inhibitor of STAT3 that inhibits phosphorylation of STAT3. LLL12 enhances the inhibitory effects of cisplatin and paclitaxel on the generation, migration, and growth of ovarian cancer cells.

-
GC67706
LY5
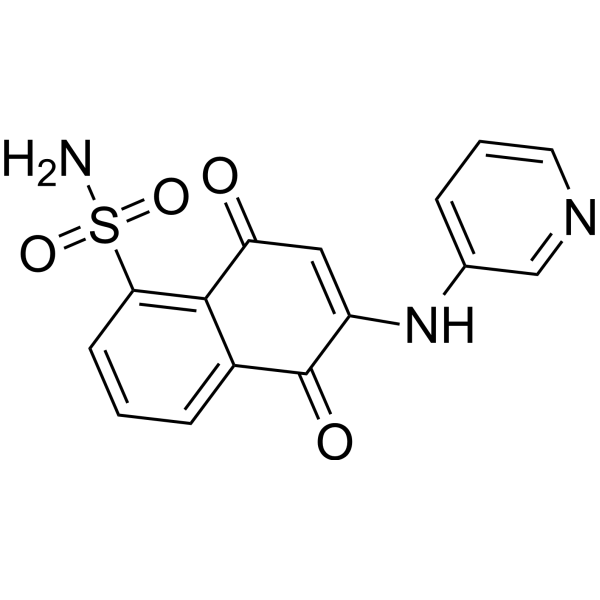
-
GC44213
ML115
CID 6619100, SID 14735210
Signal transducer and activator of transcription 3 (STAT3) is a cytokine-inducible transcription factor with roles in inflammation and cancer.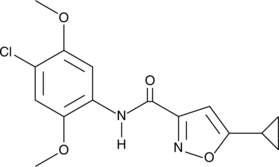
-
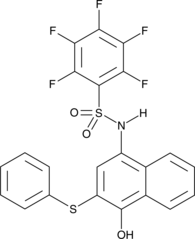
GC44227
MM-206
MM-206 is an inhibitor of STAT3 (IC50 = 1.16 μM in an SPR-based competitive assay).
-
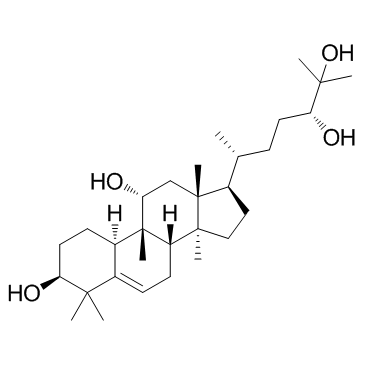
GC32789
Mogrol
Mogrol is a biometabolite of mogrosides, and acts via inhibition of the ERK1/2 and STAT3 pathways, or reducing CREB activation and activating AMPK signaling.
-
GN10436
Morusin
Mulberrochromene, NSC 649220
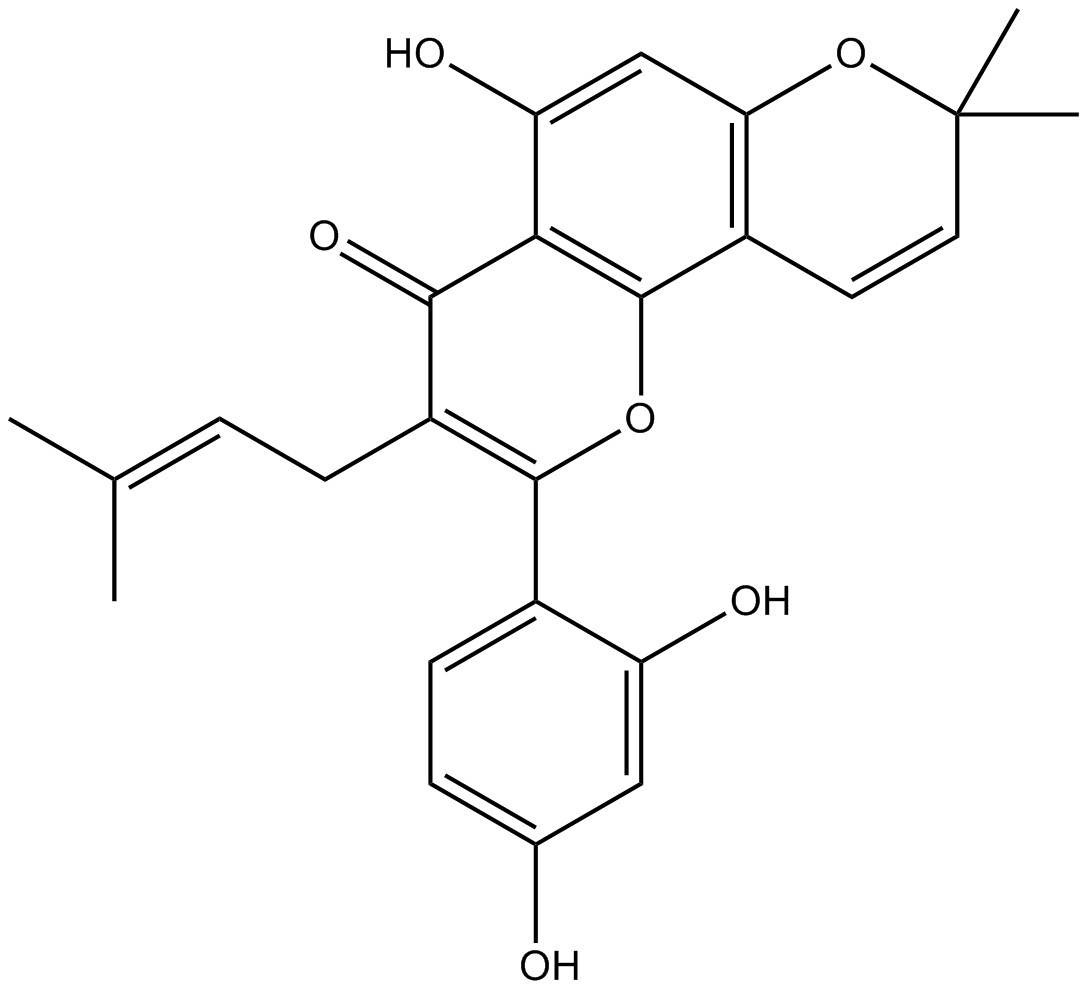
-
GC11474
Napabucasin
BBI 608
STAT3 inhibitor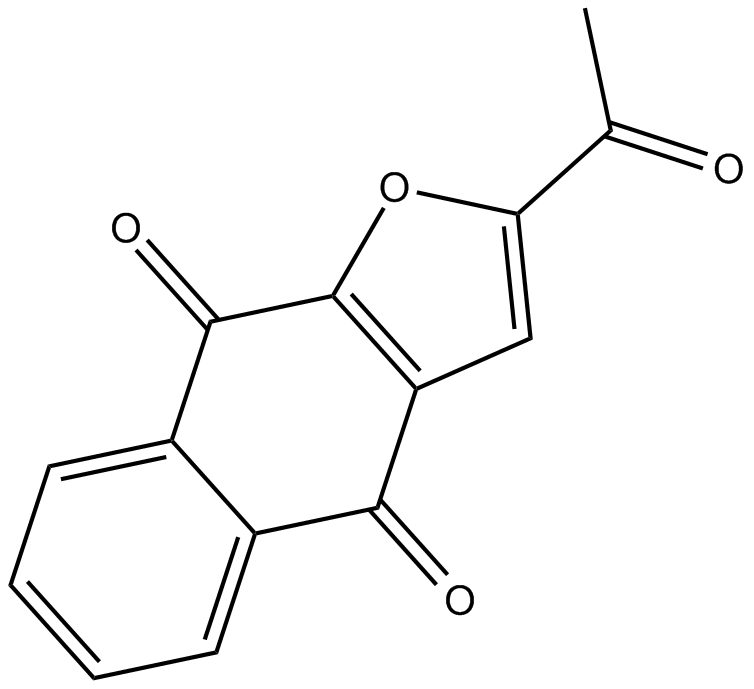
-
GC13586
Niclosamide
NSC 178296
Inhibitor of the STAT3 signaling pathway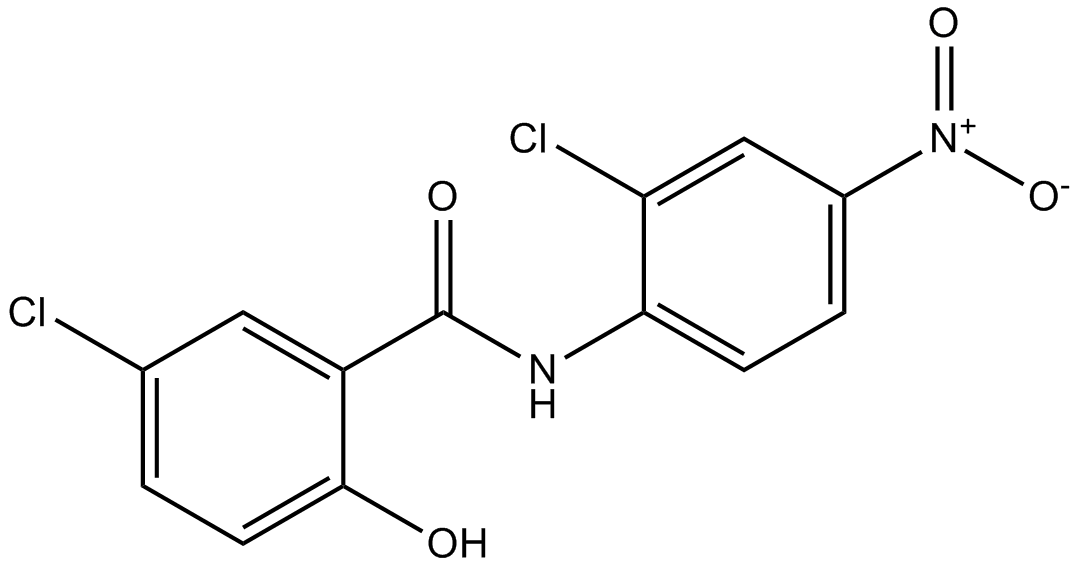
-
GC44400
Niclosamide (ethanolamine salt)
BAY-73, BAY-6076, Bayluscide, HL 2448
Niclosamide (BAY2353) olamine is an orally active antihelminthic agent used in parasitic infection research.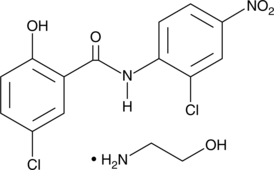
-
GC15901
Niclosamide monohydrate
agent for the treatment of most tapeworm infections
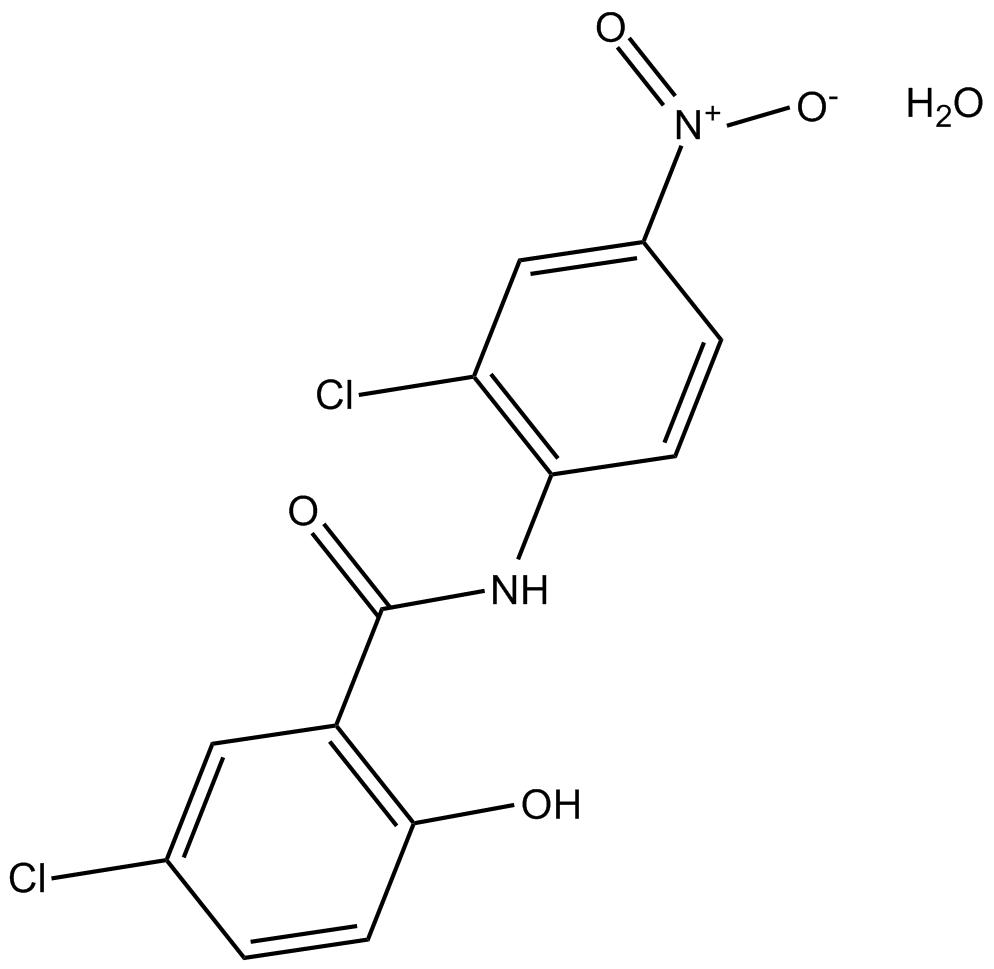
-
GC14170
Nifuroxazide
STAT inhibitor
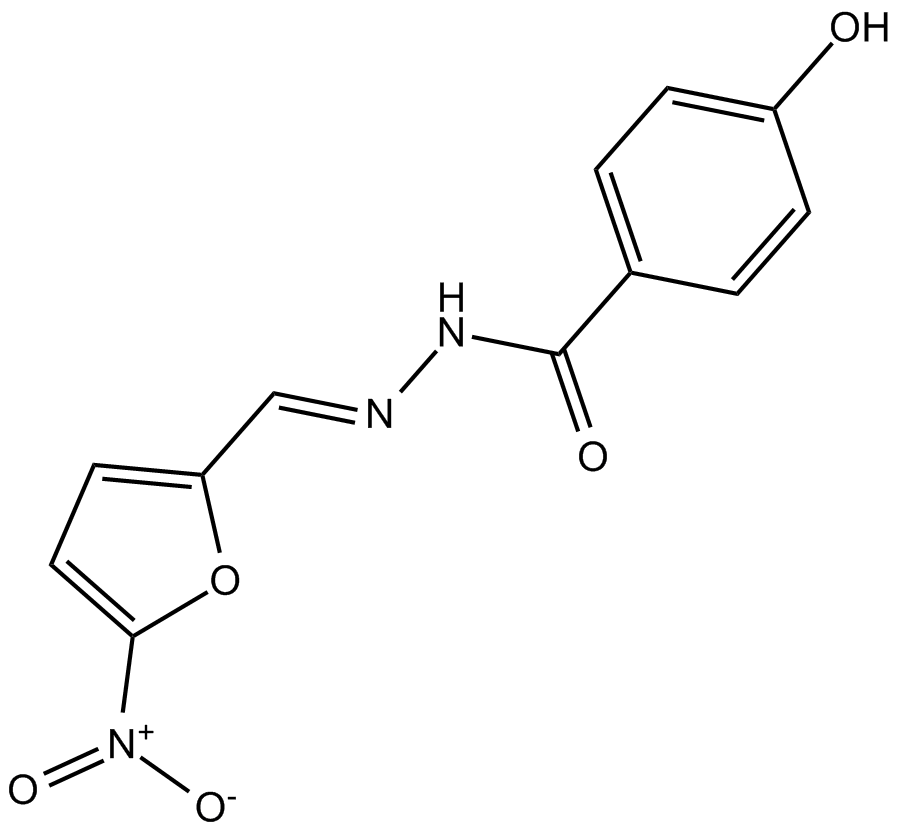
-
GC48839
Nifuroxazide-d4
An internal standard for the quantification of nifuroxazide
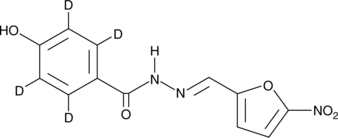
-
GC14653
NSC 74859
S3I-201;NSC74859;NSC-74859;S3I 201
Stat3 inhibitor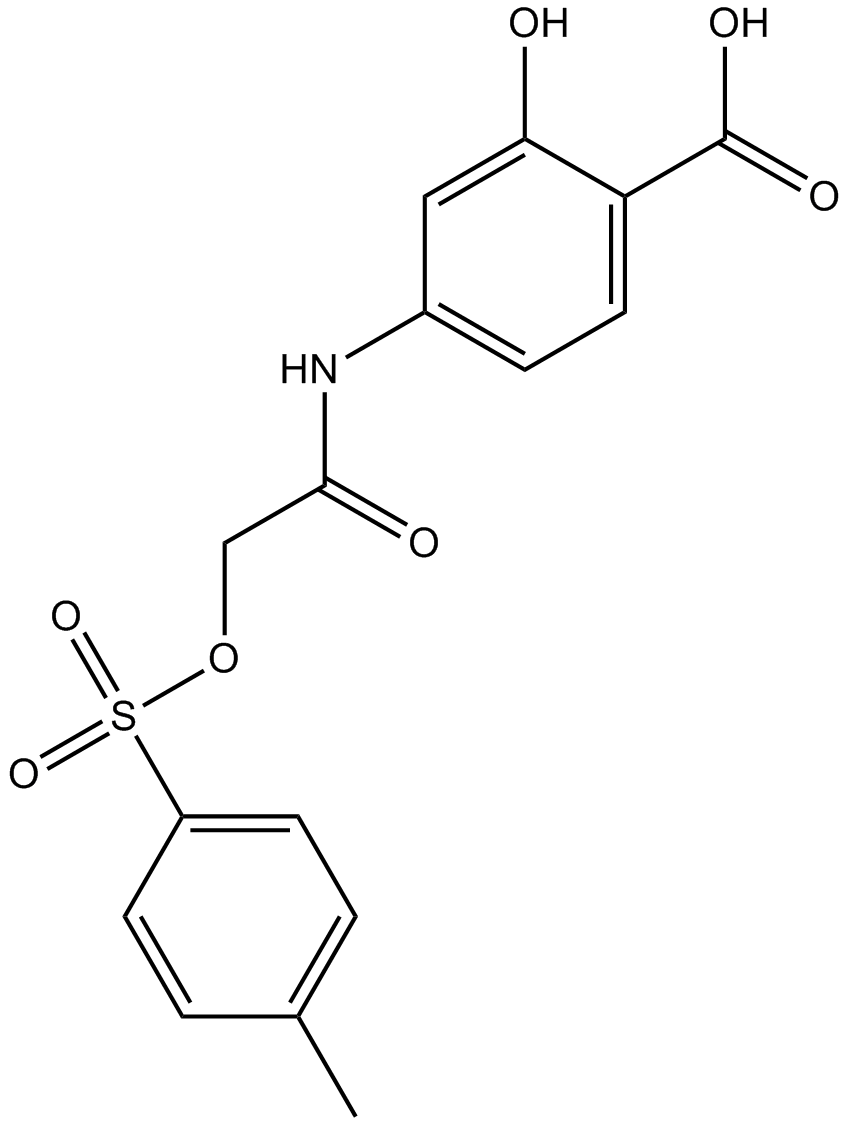
-
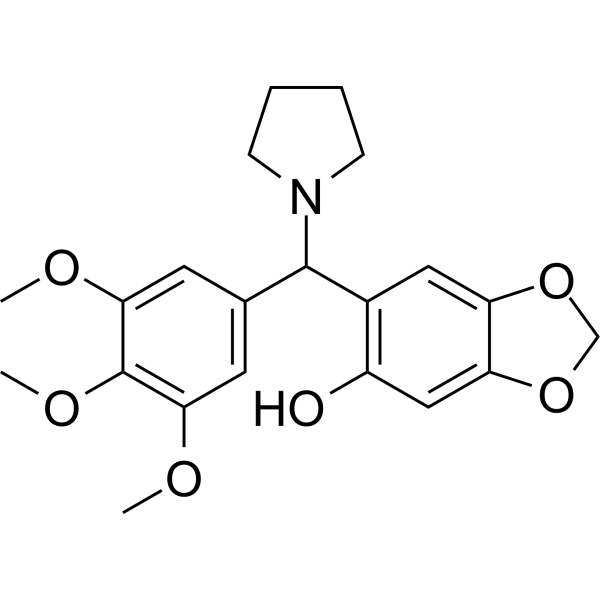
GC69594
NSC-370284
NSC-370284 is a selective inhibitor of both 10-11 translocation 1 (TET1) and 5-hydroxymethylcytosine (5hmC). It significantly inhibits TET1 expression levels by targeting STAT3/5.
-
GC12712
NT157
IRS-1/2 inhibitor, inhibits IGF-1R and STAT3 signaling pathway
-
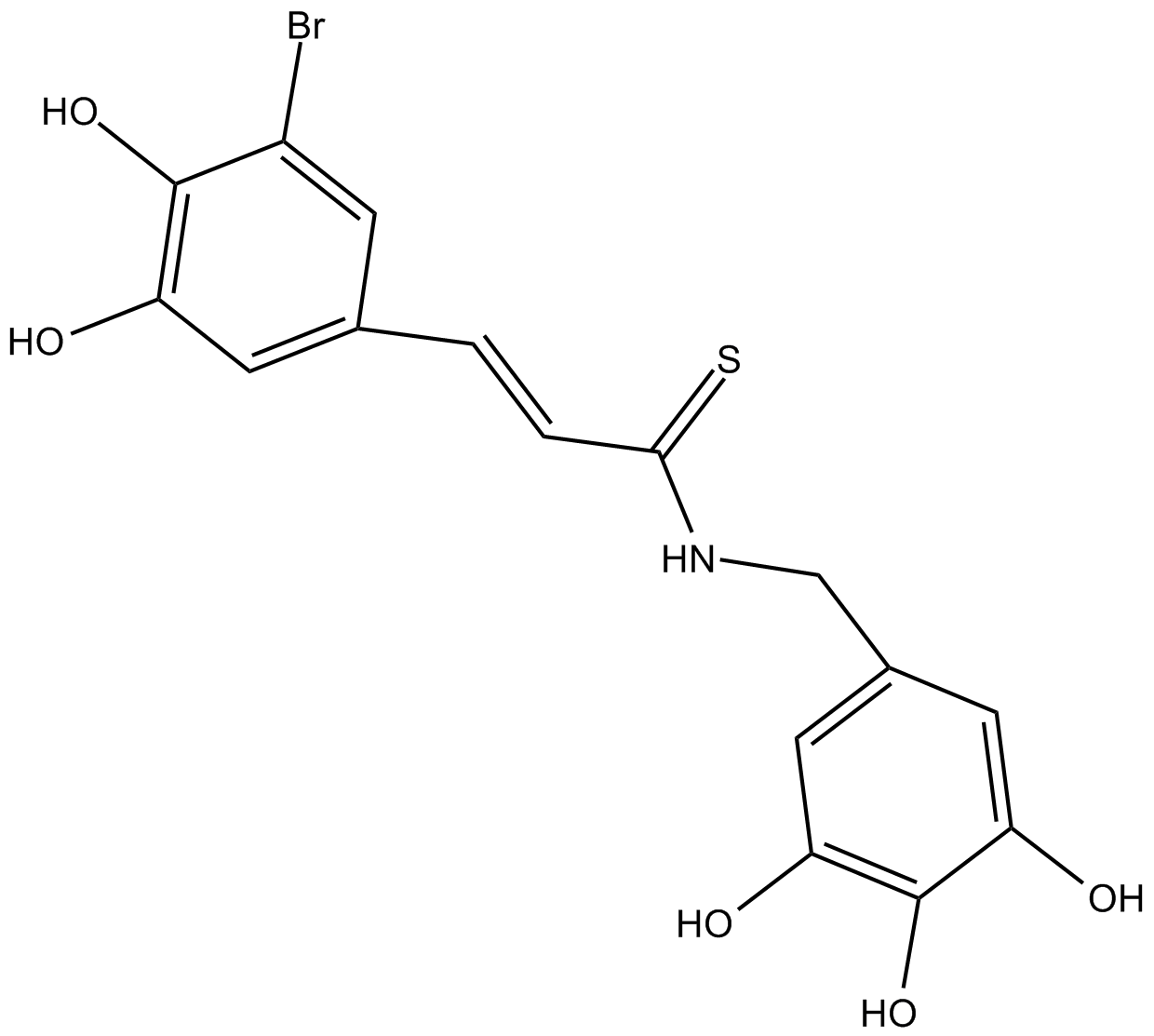
GC61627
Ochromycinone
Ochromycinone ((Rac)-STA-21) is a natural antibiotic and a STAT3 inhibitor.

-
GC69635
OSM-SMI-10B
OSM-SMI-10B is a derivative of OSM-SMI-10. Co-incubation of OSM-SMI-10B with Oncostatin M (OSM) significantly reduces STAT3 phosphorylation induced by OSM in cancer cells.

-
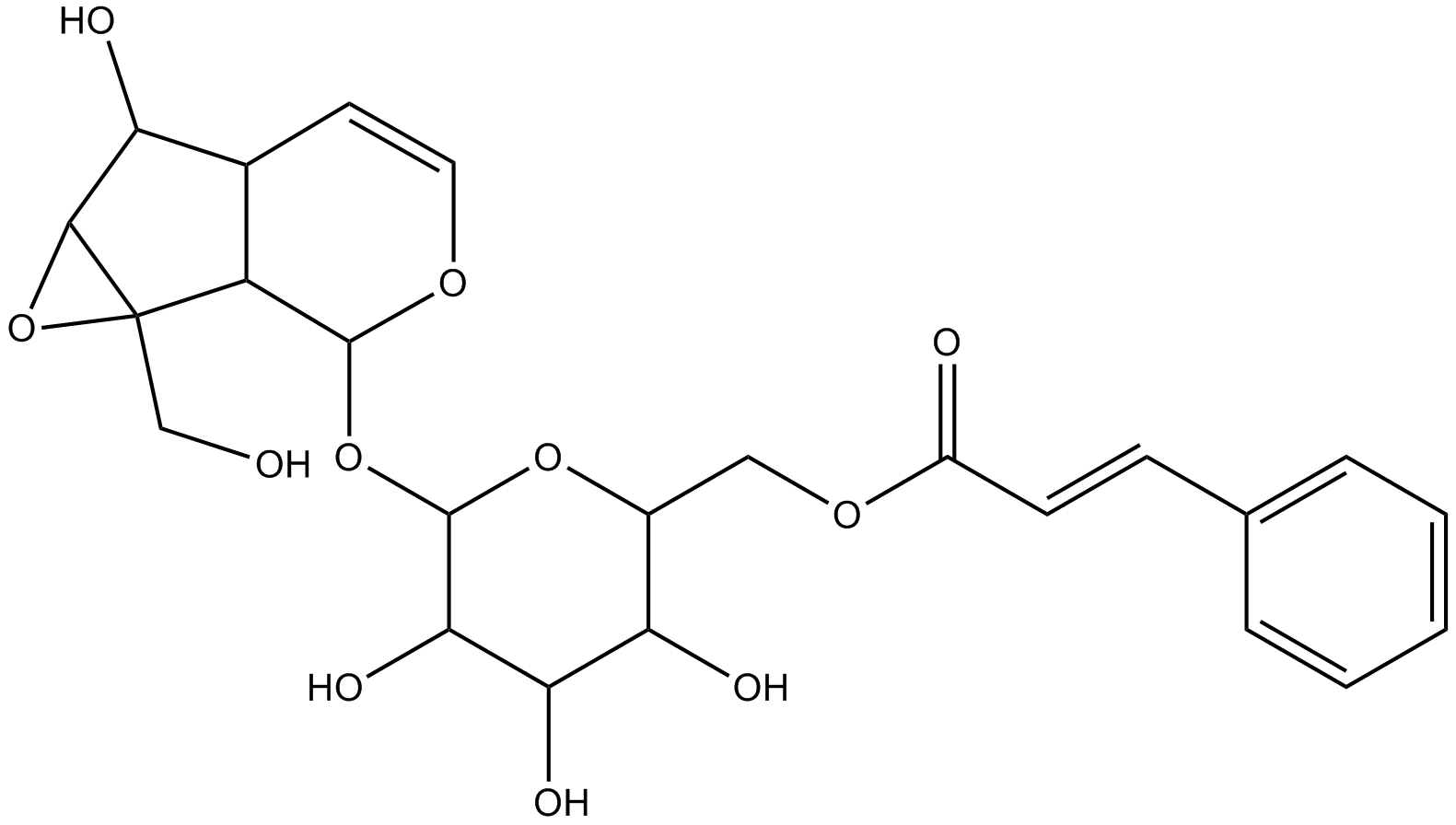
GN10314
Picroside I
6’-Cinnamoylcatalpol
-
GC10643
Pimozide
NSC 170984,Orap,R 6238
dopamine receptors inhibitor
-
GC45756
Pimozide-d4
R6238-d4
An internal standard for the quantification of pimozide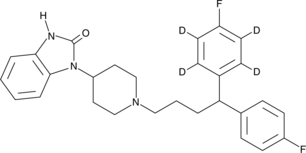
-
GC71016
PM-43I
PM-43I is a potent STAT6 inhibitor and can reduce STAT6 phosphorylation level.

-
GC71017
PM-81I
PM-81I is a potent STAT6 inhibitor (targeting the SH2 structural domain) that effectively reduces STAT6 phosphorylation levels.

-
GC34742
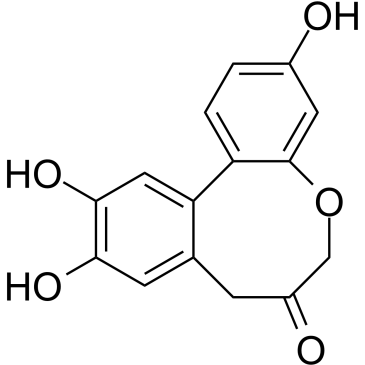
Protosappanin A
Protosappanin A (PTA), an immunosuppressive ingredient and major biphenyl compound isolated from Caesalpinia sappan L, suppresses JAK2/STAT3-dependent inflammation pathway through down-regulating the phosphorylation of JAK2 and STAT3.
-
GC39081
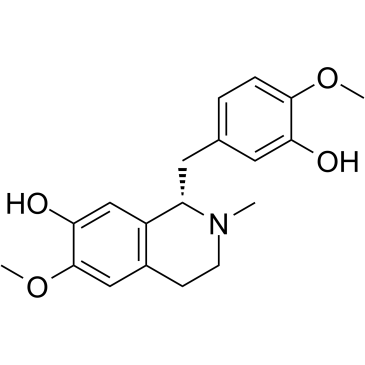
Reticuline
Reticuline shows anti-inflammatory effects through JAK2/STAT3 and NF-κB signaling pathways.
-
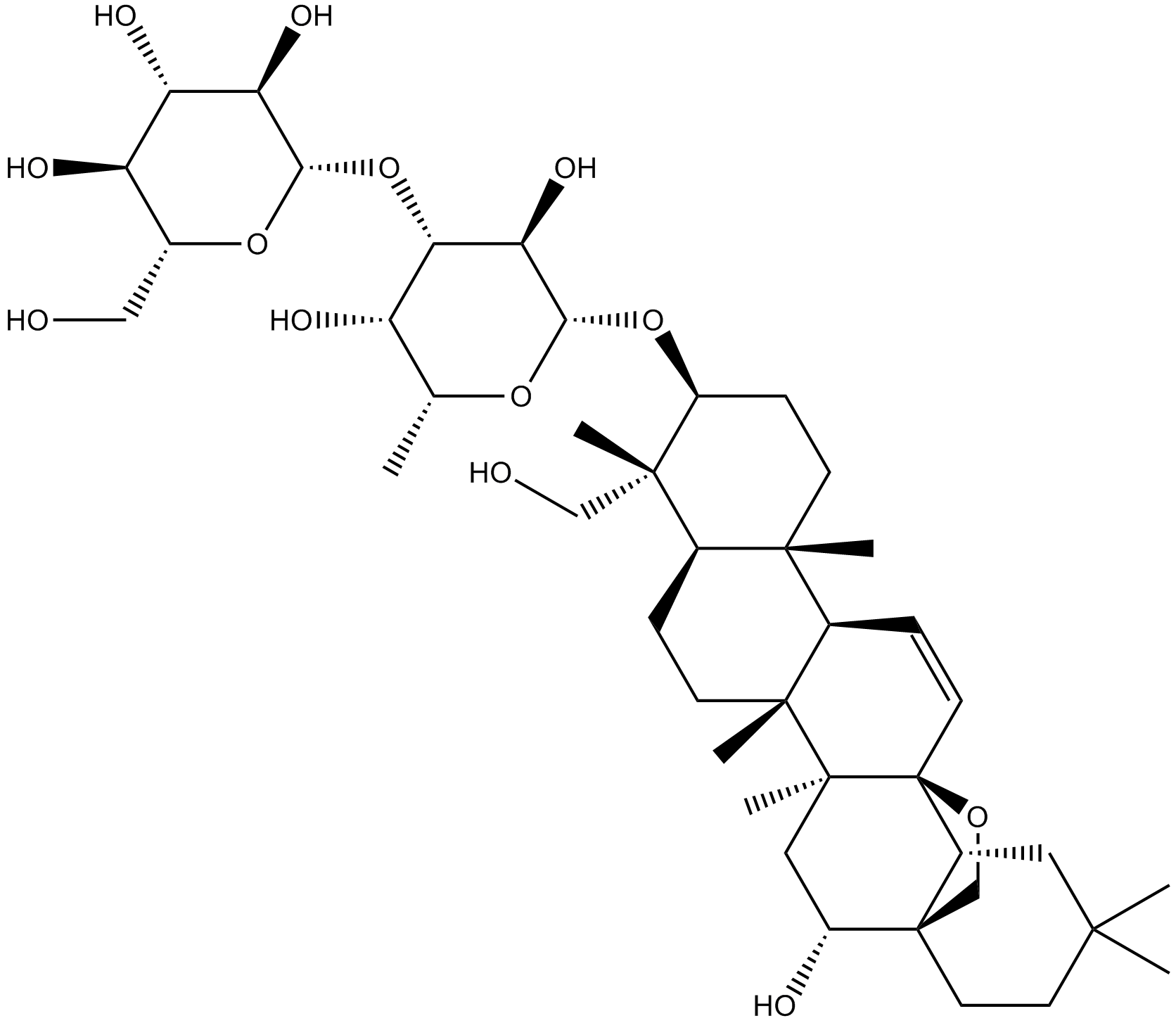
GN10164
Saikosaponin D
-
GC69854
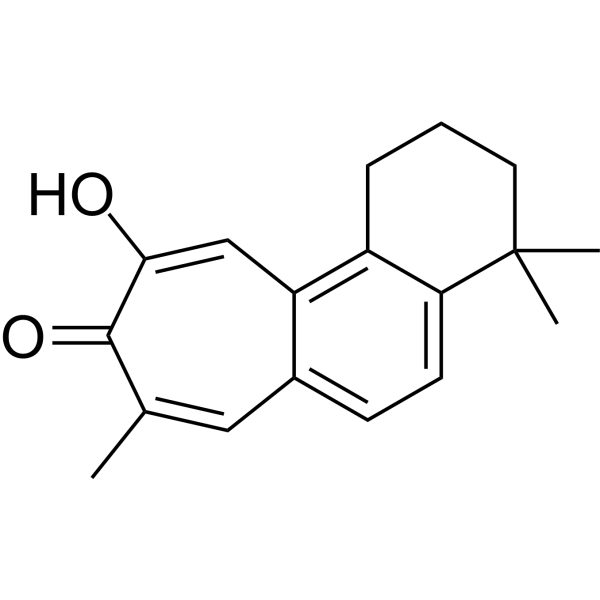
Salviolone
Salviolone is a natural diterpenoid derivative that can combat melanoma cells. Salviolone exhibits multifunctional effects on melanoma by blocking cell cycle progression, STAT3 signaling, and the malignant phenotype of A375 melanoma cells.
-
GC25899
SC-1
1-(4-Chloro-3-(trifluoromethyl)phenyl)-3-(4-(4-cyanophenoxy)phenyl)urea, STAT3-IN-7
SC-1 (1-(4-Chloro-3-(trifluoromethyl)phenyl)-3-(4-(4-cyanophenoxy)phenyl)urea, STAT3-IN-7), a Sorafenib analogue and potently inhibits the phosphorylation of STAT3, induces cell apoptosis through SHP-1 dependent STAT3 inactivation.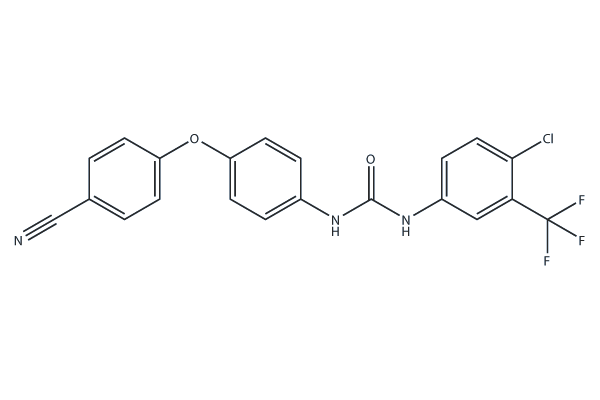
-
GC61524
SC-43
SC-43, a Sorafenib derivative, is a potent and orally active SHP-1 (PTPN6) agonist. SC-43 inhibits the phosphorylation of STAT3 and induces cell apoptosis. SC-43 has anti-fibrotic and anticancer effects.

-
GN10624
Scutellarin
Scutellarein 7-O-β-D-glucuronide
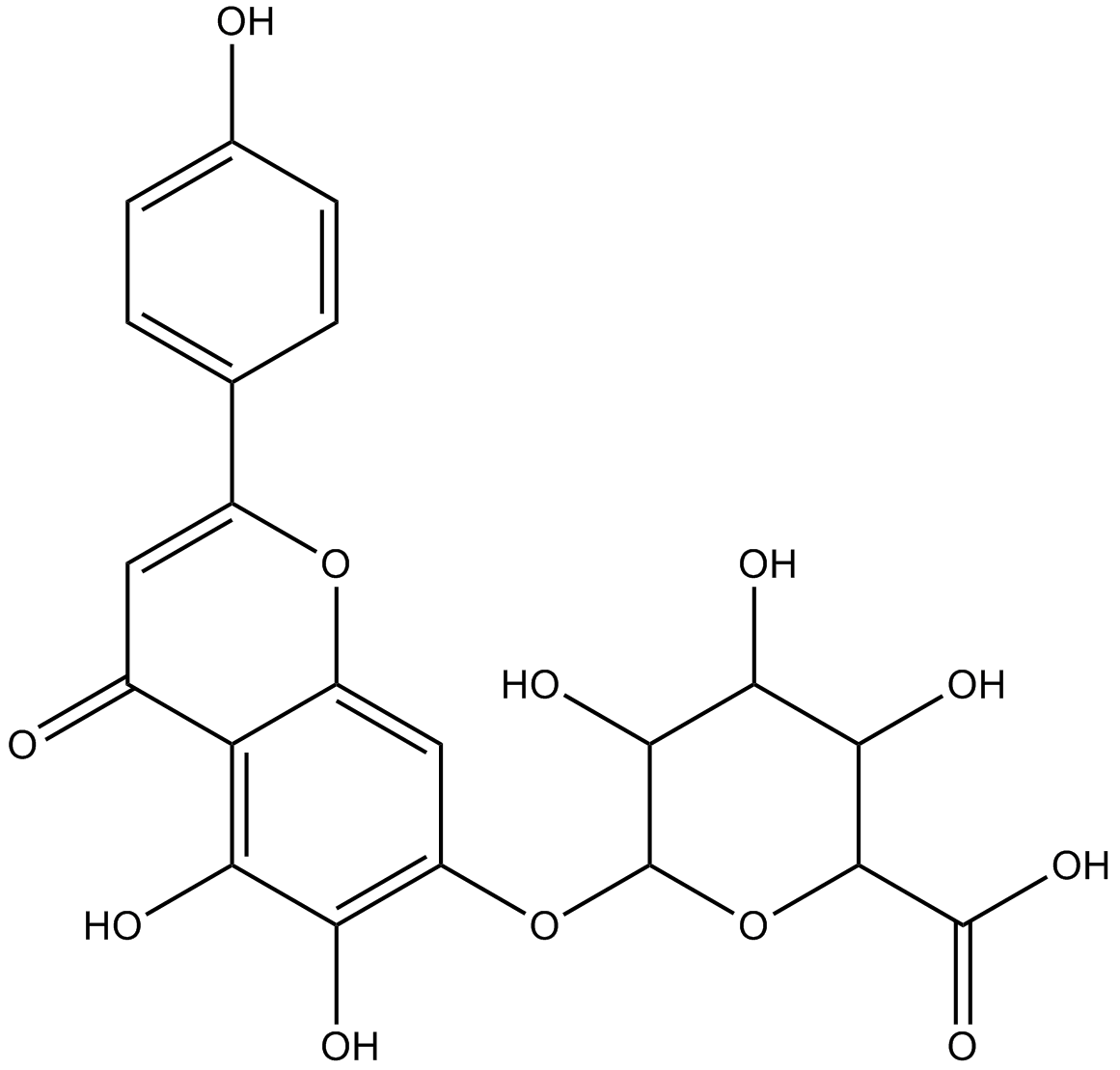
-
GC44880
SD 1029
JAK2 Inhibitor III, Janus-Associated Kinase 2 Inhibitor III
A JAK2 inhibitor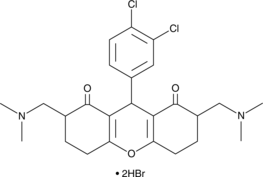
-
GC14658
SH-4-54
STAT inhibitor, potent
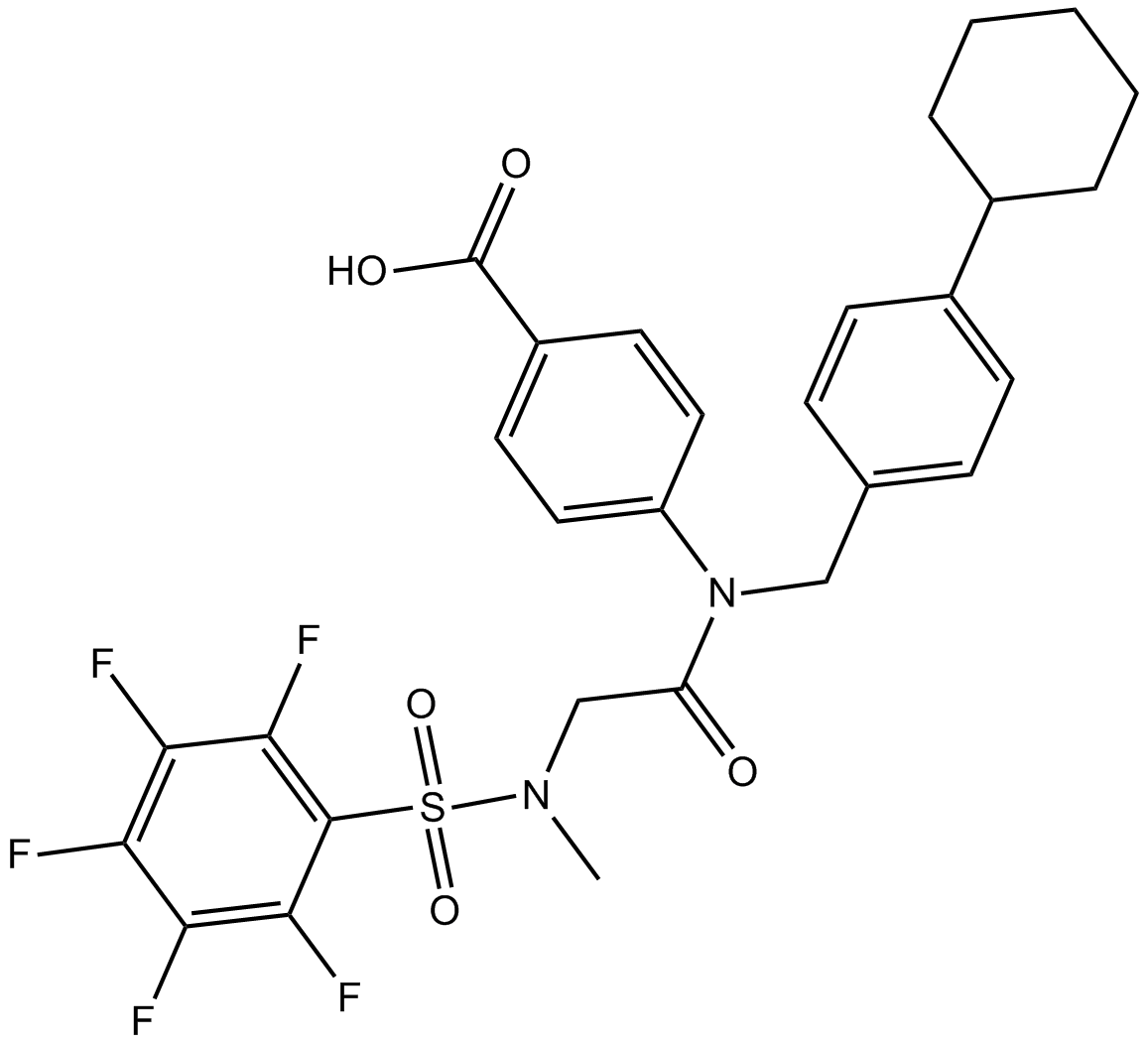
-
GC32928
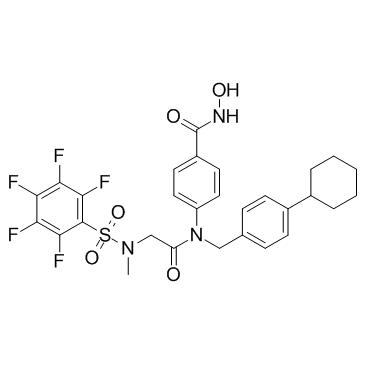
SH5-07
SH5-07 is a hydroxamic acid based Stat3 inhibitor with an IC50 of 3.9 μM in in vitro assay.
-
GC17761
STA-21
Ochromycinone
A STAT3 inhibitor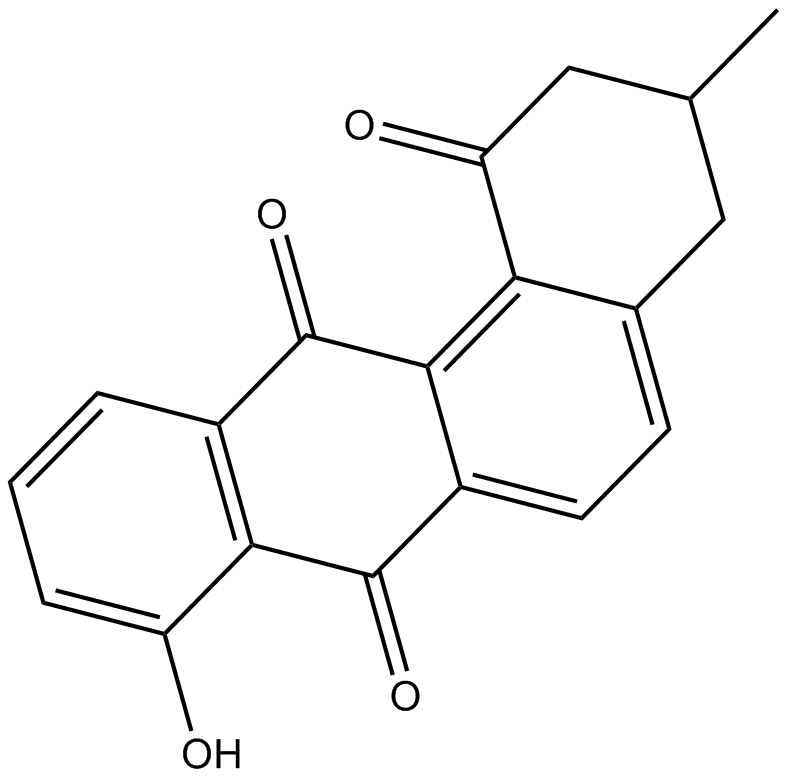
-
GC62559
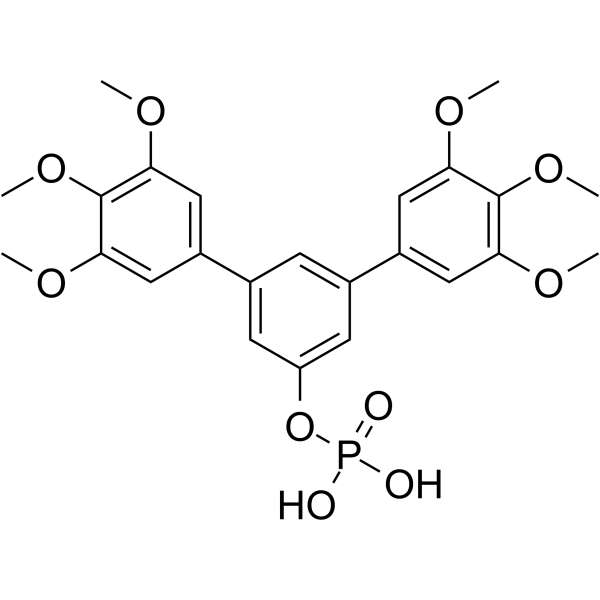
Stafia-1
Stafia-1 is a potent STAT5a inhibitor (K i=10.9 μM, IC50=22.2 μM). Stafia-1 displays high selectivity over STAT5b and other STAT family members.
-
GC63203
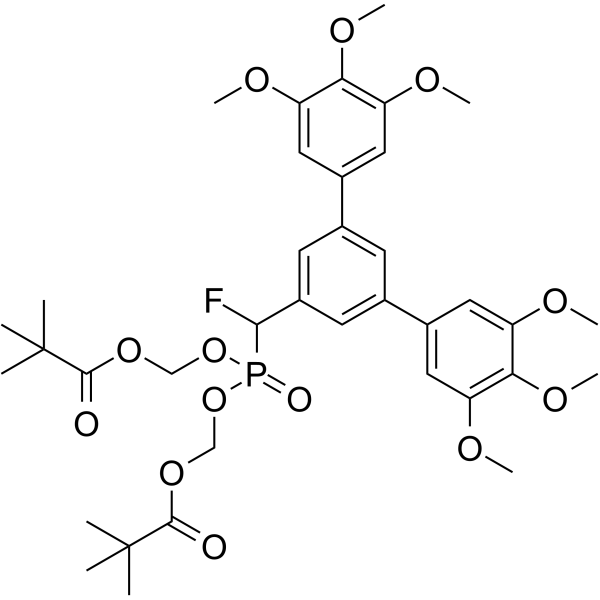
Stafia-1-dipivaloyloxymethyl ester
Stafia-1-dipivaloyloxymethyl ester (compound 27, 0-200 μM) decreases pSTAT5a expression significantly, and has no obvious inhibition on pSTAT5b.
-
GC62254
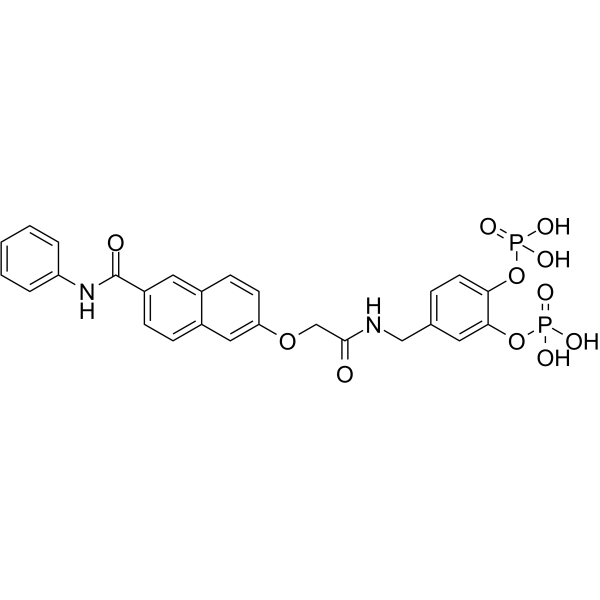
Stafib-1
Stafib-1 is the first selective inhibitor of the STAT5b SH2 domain, with a Ki of 44 nM and an IC50 of 154 nM.
-
GC63204
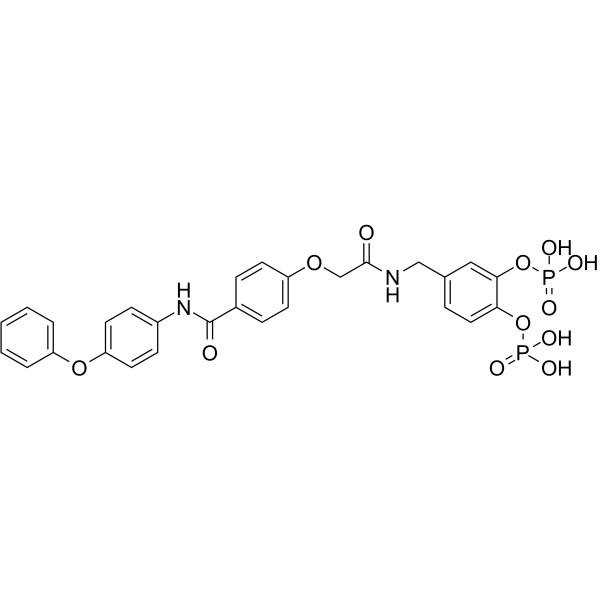
Stafib-2
Stafib-2 is a potent and selctive inhibitor of the transcription factor STAT5b, with an IC50 of 82 nM and 1.7 μM for STAT5b and STAT5a, respectively.
-
GC52293
STAT3 Inhibitor 4m
Signal Transducer and Activator of Transcription 3 Inhibitor 4m
A STAT3 inhibitor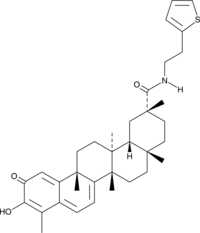
-
GC37688
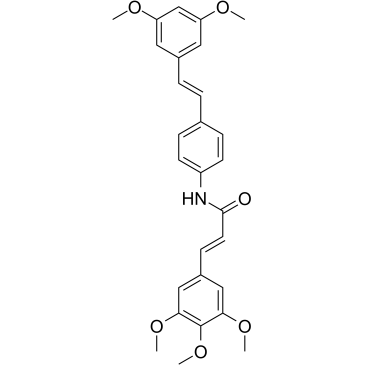
STAT3-IN-1
STAT3-IN-1 (compound 7d) is an excellent, selective and orally active STAT3 inhibitor, with IC50 values of 1.82 μM and 2.14 μM in HT29 and MDA-MB 231 cells, respectively. STAT3-IN-1 (compound 7d) induces tumor apoptosis.
-
GC69952
STAT3-IN-11
STAT3-IN-11 (7a) is a selective inhibitor of STAT3 that can inhibit the phosphorylation of the pTyr705 site on STAT3. It can also inhibit downstream genes (Survivin and Mcl-1) without affecting upstream tyrosine kinases (Src and JAK2) and p-STAT1 expression. STAT3-IN-11 can induce apoptosis in cancer cells, making it a promising candidate for the discovery of STAT3 inhibitors and anti-tumor agents.