Immunology/Inflammation
The immune and inflammation-related pathway including the Toll-like receptors pathway, the B cell receptor signaling pathway, the T cell receptor signaling pathway, etc.
Toll-like receptors (TLRs) play a central role in host cell recognition and responses to microbial pathogens. TLR4 initially recruits TIRAP and MyD88. MyD88 then recruits IRAKs, TRAF6, and the TAK1 complex, leading to early-stage activation of NF-κB and MAP kinases [1]. TLR4 is endocytosed and delivered to intracellular vesicles and forms a complex with TRAM and TRIF, which then recruits TRAF3 and the protein kinases TBK1 and IKKi. TBK1 and IKKi catalyze the phosphorylation of IRF3, leading to the expression of type I IFN [2].
BCR signaling is initiated through ligation of mIg under conditions that induce phosphorylation of the ITAMs in CD79, leading to the activation of Syk. Once Syk is activated, the BCR signal is transmitted via a series of proteins associated with the adaptor protein B-cell linker (Blnk, SLP-65). Blnk binds CD79a via non-ITAM tyrosines and is phosphorylated by Syk. Phospho-Blnk acts as a scaffold for the assembly of the other components, including Bruton’s tyrosine kinase (Btk), Vav 1, and phospholipase C-gamma 2 (PLCγ2) [3]. Following the assembly of the BCR-signalosome, GRB2 binds and activates the Ras-guanine exchange factor SOS, which in turn activates the small GTPase RAS. The original RAS signal is transmitted and amplified through the mitogen-activated protein kinase (MAPK) pathway, which including the serine/threonine-specific protein kinase RAF followed by MEK and extracellular signal related kinases ERK 1 and 2 [4]. After stimulation of BCR, CD19 is phosphorylated by Lyn. Phosphorylated CD19 activates PI3K by binding to the p85 subunit of PI3K and produce phosphatidylinositol-3,4,5-trisphosphate (PIP3) from PIP2, and PIP3 transmits signals downstream [5].
Central process of T cells responding to specific antigens is the binding of the T-cell receptor (TCR) to specific peptides bound to the major histocompatibility complex which expressed on antigen-presenting cells (APCs). Once TCR connected with its ligand, the ζ-chain–associated protein kinase 70 molecules (Zap-70) are recruited to the TCR-CD3 site and activated, resulting in an initiation of several signaling cascades. Once stimulation, Zap-70 forms complexes with several molecules including SLP-76; and a sequential protein kinase cascade is initiated, consisting of MAP kinase kinase kinase (MAP3K), MAP kinase kinase (MAPKK), and MAP kinase (MAPK) [6]. Two MAPK kinases, MKK4 and MKK7, have been reported to be the primary activators of JNK. MKK3, MKK4, and MKK6 are activators of P38 MAP kinase [7]. MAP kinase pathways are major pathways induced by TCR stimulation, and they play a key role in T-cell responses.
Phosphoinositide 3-kinase (PI3K) binds to the cytosolic domain of CD28, leading to conversion of PIP2 to PIP3, activation of PKB (Akt) and phosphoinositide-dependent kinase 1 (PDK1), and subsequent signaling transduction [8].
References
[1] Kawai T, Akira S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors[J]. Nature immunology, 2010, 11(5): 373-384.
[2] Kawai T, Akira S. Toll-like receptors and their crosstalk with other innate receptors in infection and immunity[J]. Immunity, 2011, 34(5): 637-650.
[3] Packard T A, Cambier J C. B lymphocyte antigen receptor signaling: initiation, amplification, and regulation[J]. F1000Prime Rep, 2013, 5(40.10): 12703.
[4] Zhong Y, Byrd J C, Dubovsky J A. The B-cell receptor pathway: a critical component of healthy and malignant immune biology[C]//Seminars in hematology. WB Saunders, 2014, 51(3): 206-218.
[5] Baba Y, Matsumoto M, Kurosaki T. Calcium signaling in B cells: regulation of cytosolic Ca 2+ increase and its sensor molecules, STIM1 and STIM2[J]. Molecular immunology, 2014, 62(2): 339-343.
[6] Adachi K, Davis M M. T-cell receptor ligation induces distinct signaling pathways in naive vs. antigen-experienced T cells[J]. Proceedings of the National Academy of Sciences, 2011, 108(4): 1549-1554.
[7] Rincón M, Flavell R A, Davis R A. The Jnk and P38 MAP kinase signaling pathways in T cell–mediated immune responses[J]. Free Radical Biology and Medicine, 2000, 28(9): 1328-1337.
[8] Bashour K T, Gondarenko A, Chen H, et al. CD28 and CD3 have complementary roles in T-cell traction forces[J]. Proceedings of the National Academy of Sciences, 2014, 111(6): 2241-2246.
Ziele für Immunology/Inflammation
- Cyclic GMP-AMP Synthase(2)
- Apoptosis(178)
- 5-Lipoxygenase(18)
- TLR(98)
- Papain(1)
- PGDS(1)
- PGE synthase(24)
- SIKs(10)
- IκB/IKK(60)
- AP-1(3)
- KEAP1-Nrf2(42)
- NOD1(1)
- NF-κB(224)
- Interleukin Related(147)
- 15-lipoxygenase(2)
- Others(10)
- Aryl Hydrocarbon Receptor(33)
- CD73(16)
- Complement System(52)
- Galectin(31)
- IFNAR(21)
- NO Synthase(75)
- NOD-like Receptor (NLR)(46)
- STING(86)
- Reactive Oxygen Species(402)
- FKBP(11)
- eNOS(4)
- iNOS(24)
- nNOS(20)
- Glutathione(37)
- Adaptive Immunity(144)
- Allergy(129)
- Arthritis(25)
- Autoimmunity(134)
- Gastric Disease(64)
- Immunosuppressants(27)
- Immunotherapeutics(3)
- Innate Immunity(411)
- Pulmonary Diseases(76)
- Reactive Nitrogen Species(43)
- Specialized Pro-Resolving Mediators(42)
- Reactive Sulfur Species(24)
Produkte für Immunology/Inflammation
- Bestell-Nr. Artikelname Informationen
-
GC50569
NLRP3-IN-2
NLRP3-IN-2, ein Zwischensubstrat bei der Synthese von Glyburid, hemmt die Bildung des NLRP3-Inflammasoms in Kardiomyozyten und begrenzt die InfarktgrÖße nach myokardialer IschÄmie/Reperfusion bei der Maus, ohne den Glukosestoffwechsel zu beeintrÄchtigen.

-
GC45194
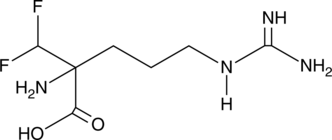
α-(difluoromethyl)-DL-Arginine
DFMA, RMI 71897
Bacteria synthesize the cellular growth factor putrescine through a number of pathways.
-
GC65446
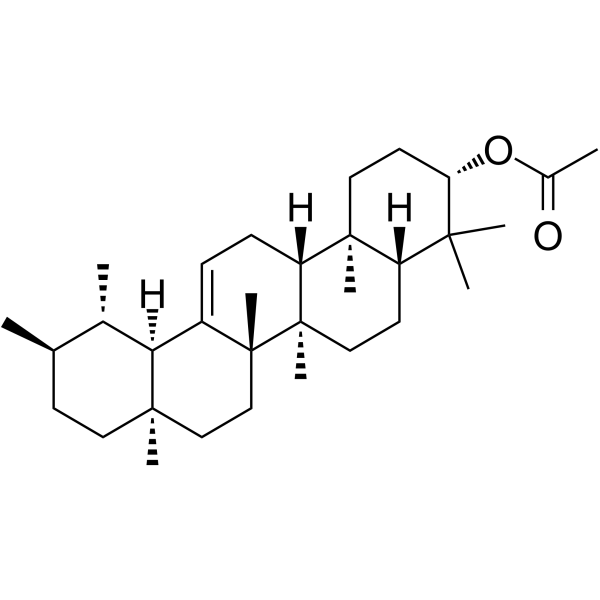
α-Amyrin acetate
α-Amyrinacetat, ein natürliches Triterpenoid, hat eine entzündungshemmende Wirkung, ein krampflösendes Profil und eine entspannende Wirkung.
-
GC49838
α-Cortolone
20α-Cortolone, NSC 59872
A metabolite of cortisol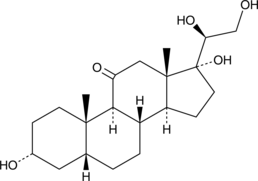
-
GC48279
α-D-Glucose-1-phosphate (sodium salt hydrate)
α-D-Glc 1-P
An intermediate in glycogen metabolism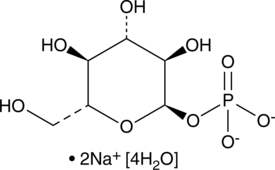
-
GC52253
α-Enolase (1-19)-biotin Peptide
Enolase-1 (1-19)-biotin
A biotinylated α-enolase peptide
-
GC45206
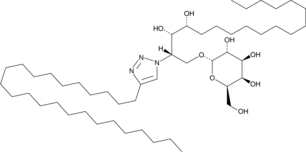
α-GalCer analog 8
α-Galactosylceramide analog 8
α-Galactosylceramide analog 8 (α-GalCer analog 8) is a triazole derivative of α-galactosylceramide.
-
GC40262
α-Humulene
αCaryophyllene, (±)-αHumulene
α-Humulen ist ein Hauptbestandteil von Tanacetum vulgare L.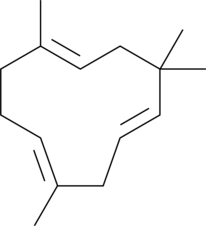
-
GC45601
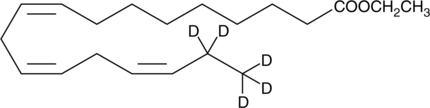
α-Linolenic Acid ethyl ester-d5
ALAEE-d5, Ethyl α-Linolenate-d5, Ethyl Linolenate-d5, LAEE-d5, Linolenic Acid ethyl ester-d5
-
GC48292
α-MSH (human, mouse, rat, porcine, bovine, ovine) (trifluoroacetate salt)
α-Melanocyte-stimulating Hormone, Ac-SYSMEHFRWGKPV-NH2
α-MSH (α-Melanozyten-stimulierendes Hormon) TFA, ein endogenes Neuropeptid, ist ein endogener Melanocortinrezeptor 4 (MC4R)-Agonist mit entzÜndungshemmenden und antipyretischen AktivitÄten.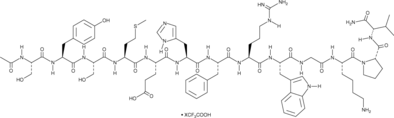
-
GC41499
α-Phellandrene
p-Mentha-1,5-diene, (±)-α-Phellandrene
α-Phellandrene is a cyclic monoterpene that has been found in various plants, including Cannabis, and has diverse biological activities.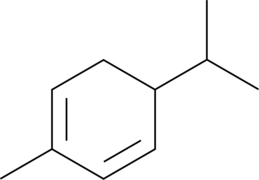
-
GC63941
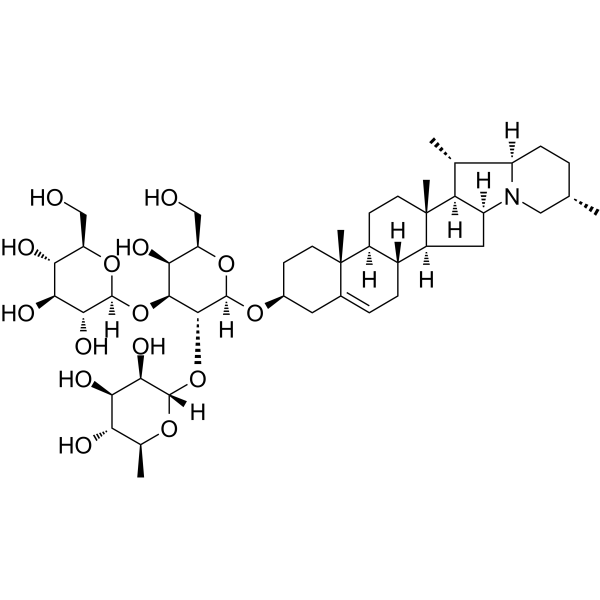
α-Solanine
α-Solanin, eine bioaktive Komponente und eines der wichtigsten steroidalen Glykoalkaloide in Kartoffeln, hemmt nachweislich das Wachstum und induziert Apoptose in Krebszellen.
-
GC67618
α-Tocopherol phosphate disodium
alpha-Tocopherol phosphate disodium; TocP disodium; Vitamin E phosphate disodium
α-Tocopherolphosphat (Alpha-Tocopherolphosphat)-Dinatrium, ein vielversprechendes Antioxidans, kann vor langwelligem UVA1-induziertem Zelltod schützen und UVA1-induzierte ROS in einem Hautzellmodell abfangen. α-Tocopherolphosphat-Dinatrium besitzt therapeutisches Potenzial bei der Hemmung der Apoptose und erhöht die Migrationskapazität von endothelialen Vorläuferzellen unter Bedingungen mit hohem Glukosegehalt/Hypoxie und fördert die Angiogenese.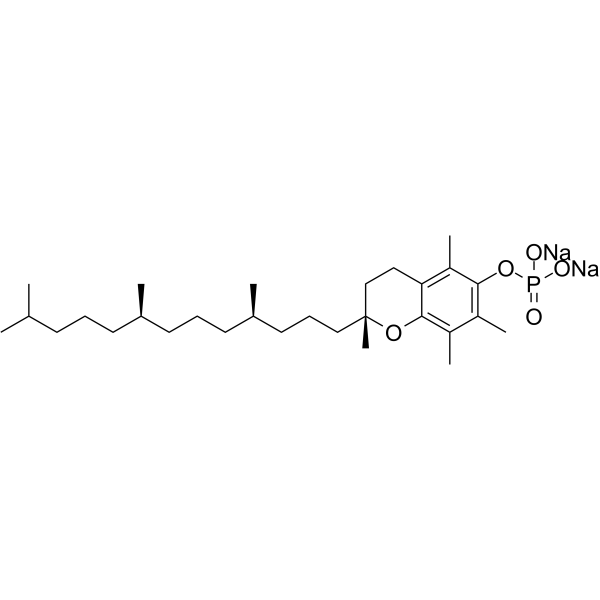
-
GC70953
α7 nAchR-JAK2-STAT3 agonist 1
α7 nAchR-JAK2-STAT3 agonist 1 is a potent α7 nAchR-JAK2-STAT3 agonist, with an IC50 value of 0.32 μM for nitric oxide (NO).

-
GC49467
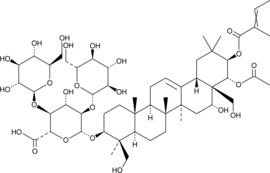
β-Aescin
A triterpenoid saponin with diverse biological activities
-
GC70787
β-Aminoarteether
β-Aminoarteether (SM934 free base) is an Artemisinin derivative with orally active.

-
GC37999
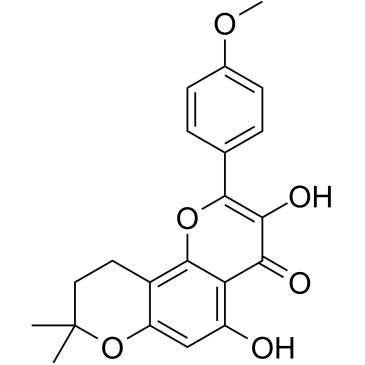
β-Anhydroicaritin
β-Anhydroicaritin wird aus Boswellia carterii Birdware isoliert, hat wichtige biologische und pharmakologische Wirkungen wie Antiosteoporose, Östrogenregulierung und Antitumoreigenschaften.
-
GC45225
β-Apooxytetracycline
β-Apo-Oxytetracycline, β-Apoterramycin
β-Apooxytetracycline is a potential impurity found in commercial preparations of oxytetracycline.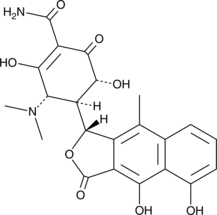
-
GC48920
β-Carboline-1-carboxylic Acid
1-Formic Acid-β-carboline
An alkaloid with diverse biological activities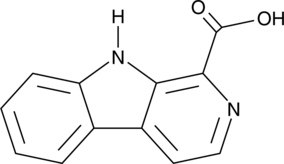
-
GC66870
β-D-Glucan
β-D-Glucan ist ein natürliches, unverdauliches Polysaccharid mit hoher Biokompatibilität, das selektiv von Erkennungsrezeptoren wie Dectin-1 und Toll-like-Rezeptoren erkannt werden kann und leicht von murinen oder menschlichen Makrophagen internalisiert werden kann, was wahrscheinlich der Fall ist Attribut zu einer Zielzustellung. β-d-Glucan ist ein enterisches Transportmittel für Probiotika.

-
GC48998
β-Defensin-1 (human) (trifluoroacetate salt)
hBD-1
An antimicrobial peptide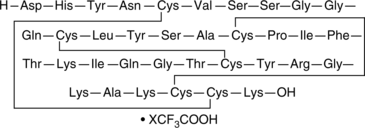
-
GC48298
β-Defensin-2 (human) (trifluoroacetate salt)
hBD-2
An antimicrobial peptide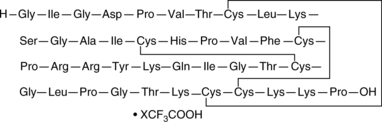
-
GC45230
β-Defensin-3 (human) (trifluoroacetate salt)
hBD-3
β-Defensin-3 is a peptide with antimicrobial properties that protects the skin and mucosal membranes of the respiratory, genitourinary, and gastrointestinal tracts.
-
GC45231
β-Defensin-4 (human) (trifluoroacetate salt)
hBD-4 (human)
β-Defensin-4 is a peptide with antimicrobial properties that protects the skin and mucosal membranes of the respiratory, genitourinary, and gastrointestinal tracts.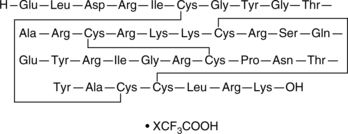
-
GC41623
β-Elemonic Acid
Elemadienonic Acid, 3-Oxotirucallenoic Acid, 3-oxo Tirucallic Acid
β-Elemonic Acid ist ein aus Boswellia papyrifera isoliertes Triterpen.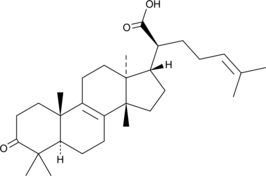
-
GC49769
β-Glucogallin
1-O-Galloyl-β-D-glucose
A plant metabolite and an aldose reductase 2 inhibitor
-
GC64619
β-Ionone
β-Ionone ist wirksam bei der Induktion von Apoptose in Adenokarzinomzellen des Magens SGC7901. Anti-Krebs-AktivitÄt.
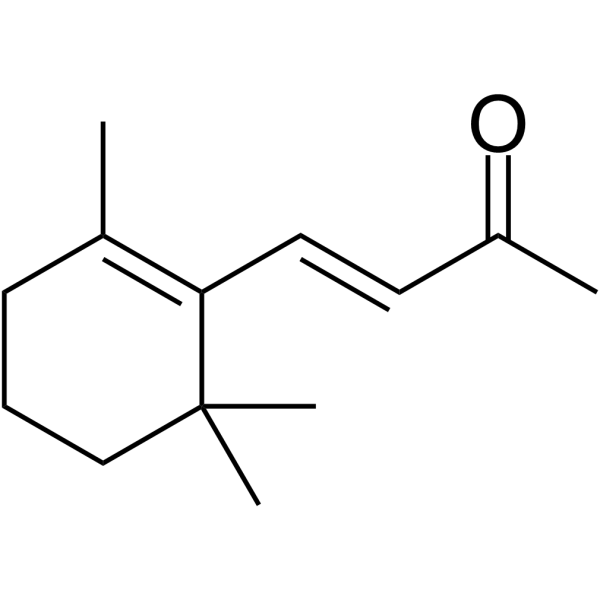
-
GC41502
β-Myrcene
NSC 406264
β-Myrcen (β-β-Myrcen), eine aromatische flüchtige Verbindung, unterdrückt die TNFα-induzierte NF-κB-Aktivität.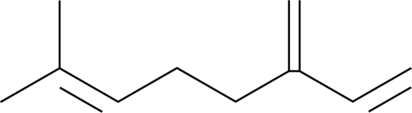
-
GC45604
β-Rubromycin
β-Rubromycin ist ein potenter und selektiver Inhibitor der RNA-gerichteten DNA-Polymerasen (reverse Transkriptase) des humanen ImmunschwÄchevirus-1 (HIV-1).
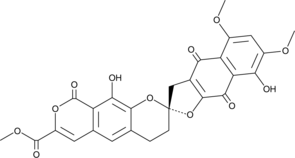
-
GC52400
γ-Glu-Ala (trifluoroacetate salt)
γ-Glutamylalanine, γ-L-Glutamyl-L-alanine
A dipeptide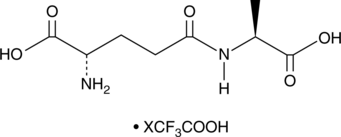
-
GC48312
γ-Glu-Cys (ammonium salt)
γ-Glutamylcysteine
An intermediate in GSH synthesis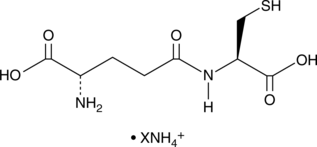
-
GC45238
δ14-Triamcinolone acetonide
14,15-dehydro Triamcinolone acetonide, Triamcinolone acetonide Impurity B
δ14-Triamcinolone acetonide is a potential impurity found in commercial preparations of triamcinolone acetonide.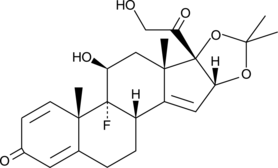
-
GC40307
δ2-cis-Hexadecenoic Acid
One of the first organisms in which quorum sensing was observed were Myxobacteria, a group of gram-negative bacteria, found mainly in soil and also common to marine and freshwater systems.
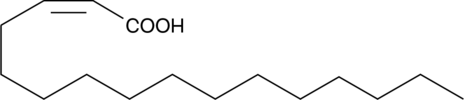
-
GC41393
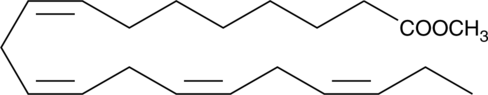
ω-3 Arachidonic Acid methyl ester
ω-3 Fatty acids, represented primarily by docosahexaenoic acid, eicosapentaenoic acid, and α-linoleate, are essential dietary nutrients required for normal growth and development.
-
GC45713
(±)-α-Tocopherol Acetate
all-rac-α-Tocopherol Acetate, DL-α-Tocopherol Acetate, DL-Vitamin E acetate
(±)-α-Tocopherolacetat ((±)-Vitamin E-Acetat), ist eine oral aktive synthetische Form von Vitamin E.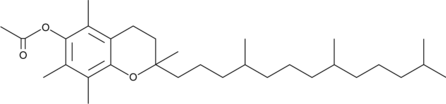
-
GC67191
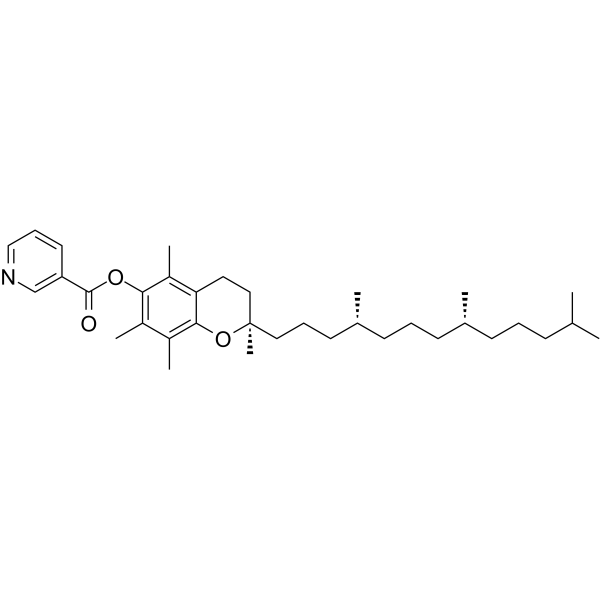
(±)-α-Tocopherol nicotinate
(±)-α-Tocopherolnicotinat, Vitamin E-Nicotinat, ist ein oral wirksames fettlösliches Antioxidans, das die Lipidperoxidation in Zellmembranen verhindert. (±)-α-Tocopherolnicotinat wird im Blut zu α hydrolysiert; -Tocopherol und Niacin und kann in Studien verwandter Gefäßerkrankungen verwendet werden.
-
GC52010
(±)-10-hydroxy-12(Z),15(Z)-Octadecadienoic Acid
αHYA, (±)-10-hydroxy-12(Z),15(Z)-ODE
An oxylipin gut microbiota metabolite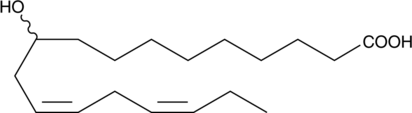
-
GC52013
(±)-10-hydroxy-12(Z)-Octadecenoic Acid
10-hydroxy-cis-12-Octadecenoic Acid
An oxylipin and metabolite of linoleic acid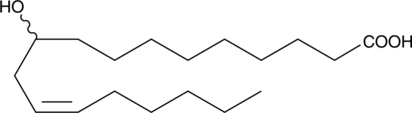
-
GC52421
(±)-10-hydroxy-12(Z)-Octadecenoic Acid-d5
10-hydroxy-cis-12-Octadecenoic Acid-d5
An internal standard for the quantification of (±)-10-hydroxy-12(Z)-octadecenoic acid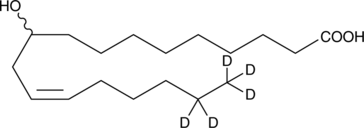
-
GC40112
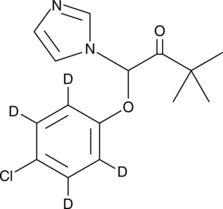
(±)-Climbazole-d4
(±)-Climbazole-d4 is intended for use as an internal standard for the quantification of climbazole by GC- or LC-MS.
-
GC50708
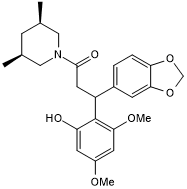
(±)-ML 209
An RORγt antagonist
-
GC39271
(±)-Naringenin
SDihydrogenistein, NSC 11855, NSC 34875, Salipurol
(±)-Naringenin ist ein natürlich vorkommendes Flavonoid.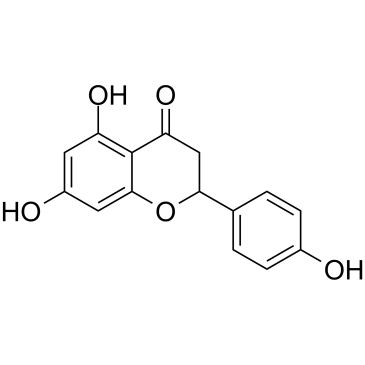
-
GC41212
(±)10(11)-EpDPA
(±)10,11-EDP, (±)10,11-EpDPE, (±)10,11-epoxy DPA, (±)10,11-epoxy Docosapentaenoic Acid
Cytochrome P450 metabolism of polyunsaturated fatty acids produces numerous bioactive epoxide regioisomers.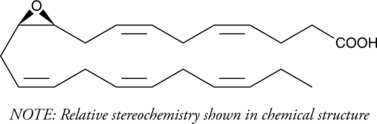
-
GC40466
(±)11(12)-EET
(±)11,12-EpETrE
(±)11(12)-EET ist ein NLRP3-Inflammasom-Inhibitor.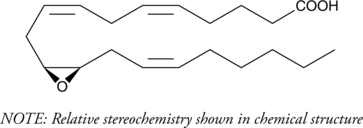
-
GC40467
(±)11-HETE
(±)11-Hydroxyeicosatetraenoic Acid
(±)11-HETE is one of the six monohydroxy fatty acids produced by the non-enzymatic oxidation of arachidonic acid.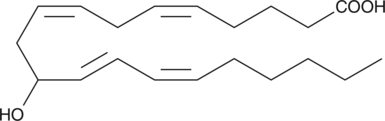
-
GC40802
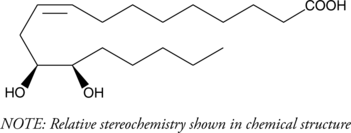
(±)12(13)-DiHOME
Isoleukotoxin diol
(±)12(13)-DiHOME is the diol form of (±)12(13)-EpOME, a cytochrome P450-derived epoxide of linoleic acid also known as isoleukotoxin.
-
GC41191
(±)13(14)-EpDPA
(±)13,14-EDP, (±)13,14-EpDPE, (±)13,14-epoxy DPA, (±)13,14-epoxy Docosapentaenoic Acid
(±)13(14)-EpDPA (13,14-EpDPE) ist das Produkt der Reaktion von Cytochrom-P-450-Epoxygenase mit Docosahexaensäure (DHA).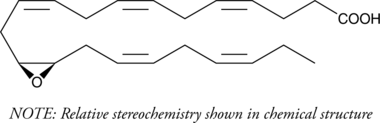
-
GC40355
(±)13-HpODE
13-Hydroperoxylinoleic acid; Linoleic acid 13-hydroperoxide
(±)13-HpODE (13-Hydroperoxylinolsäure) ist ein racemisches Gemisch von Hydroperoxiden, das durch die Oxidation von Linolsäure durch Lipoxygenase hergestellt wird.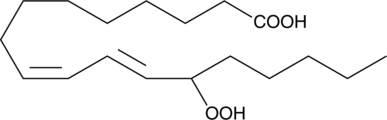
-
GC41288
(±)17(18)-EpETE-Ethanolamide
17,18-EEQ-EA, (±)17,18-EEQ-Ethanolamide, (±)17(18)-EpETE-EA, 17,18-epoxy-Eicosatetraenoic Acid Ethanolamide
(±)17(18)-EpETE-Ethanolamide is an ω-3 endocannabinoid epoxide.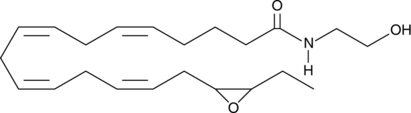
-
GC40362
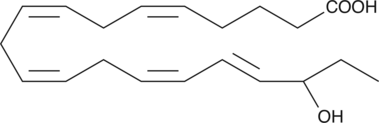
(±)18-HEPE
(±)18-HEPE is produced by non-enzymatic oxidation of EPA.
-
GC41655
(±)19(20)-EDP Ethanolamide
19,20-DHEA epoxide, 19,20-epoxy Docosapentaenoic Acid Ethanolamide, 19,20-EDP-EA, 19,20-EDP epoxide
(±)19(20)-EDP ethanolamide is an ω-3 endocannabinoid epoxide and cannabinoid (CB) receptor agonist (EC50s = 108 and 280 nM for CB1 and CB2, respectively).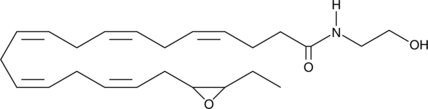
-
GC40270
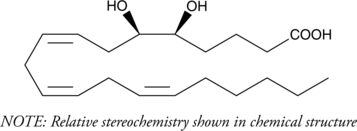
(±)5(6)-DiHET
(±)5,6-DiHETrE
5(6)-DiHET is a fully racemic version of the enantiomeric forms biosynthesized from 5(6)-EET by epoxide hydrolases.
-
GC41203
(±)7(8)-EpDPA
(±)7,8-EDP, (±)7,8-EpDPE, (±)7,8-epoxy DPA, (±)7,8-epoxy Docosapentaenoic Acid
Docosahexaenoic acid is the most abundant ω-3 fatty acid in neural tissues, especially in the brain and retina.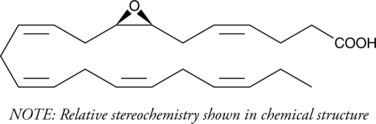
-
GC40801
(±)9(10)-DiHOME
Leukotoxin diol
(±)9(10)-DiHOME ist das Racemat von 9,10-DiHOME.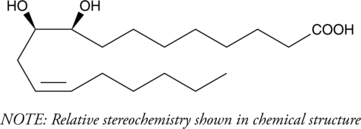
-
GC46000
(•)-Drimenol
NSC 169775
A sesquiterpene alcohol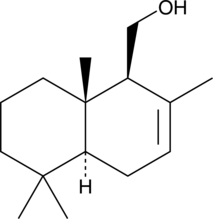
-
GC40809
(+)-β-Citronellol
(R)-Citronellol, (+)-Citronellol, (+)-(R)-Citronellol, (R)-(+)-β-Citronellol
(+)-β-Citronellol (D-Citronellol) ist ein alkoholisches Monoterpen, das in Ätherischem GeranienÖl vorkommt.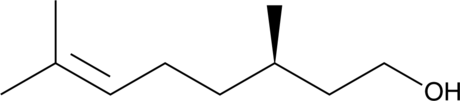
-
GC49268
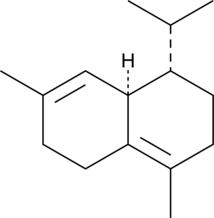
(+)-δ-Cadinene
A sesquiterpene with antimicrobial and anticancer activities
-
GC45263
(+)-D-threo-PDMP (hydrochloride)
D-PDMP
(+)-D-threo-PDMP is a ceramide analog and is one of the four possible stereoisomers of PDMP.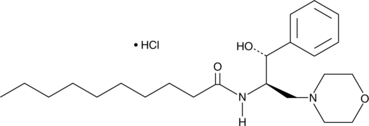
-
GC31691
(+)-DHMEQ
(1R,2R,6R)-Dehydroxymethylepoxyquinomicin; (1R,2R,6R)-DHMEQ
(+)-DHMEQ ist ein Aktivator des antioxidativen Transkriptionsfaktors Nrf2.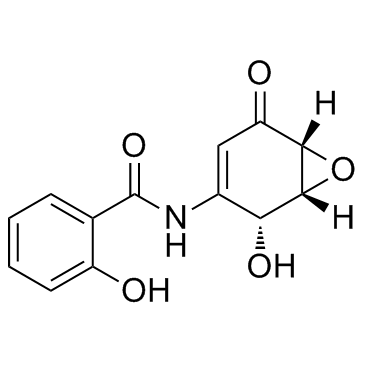
-
GC45266
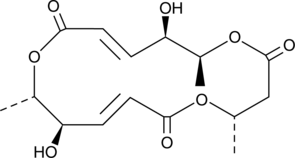
(+)-Macrosphelide A
(+)-Macrosphelide A ist ein Makrolid-Antibiotikum.
-
GC40266
(+)-Praeruptorin A
(+)-Praeruptorin A ist ein bioaktiver Hauptbestandteil von Peucedanum praeruptorum (auch bekannt als Bai-Hua Qian Hu).

-
GC18749
(+)-Rugulosin
NSC 160880, NSC 249990, Rugulosin A
(+)-Rugulosin ist ein kristalliner Farbstoff von Penicillium rugulosum Thom.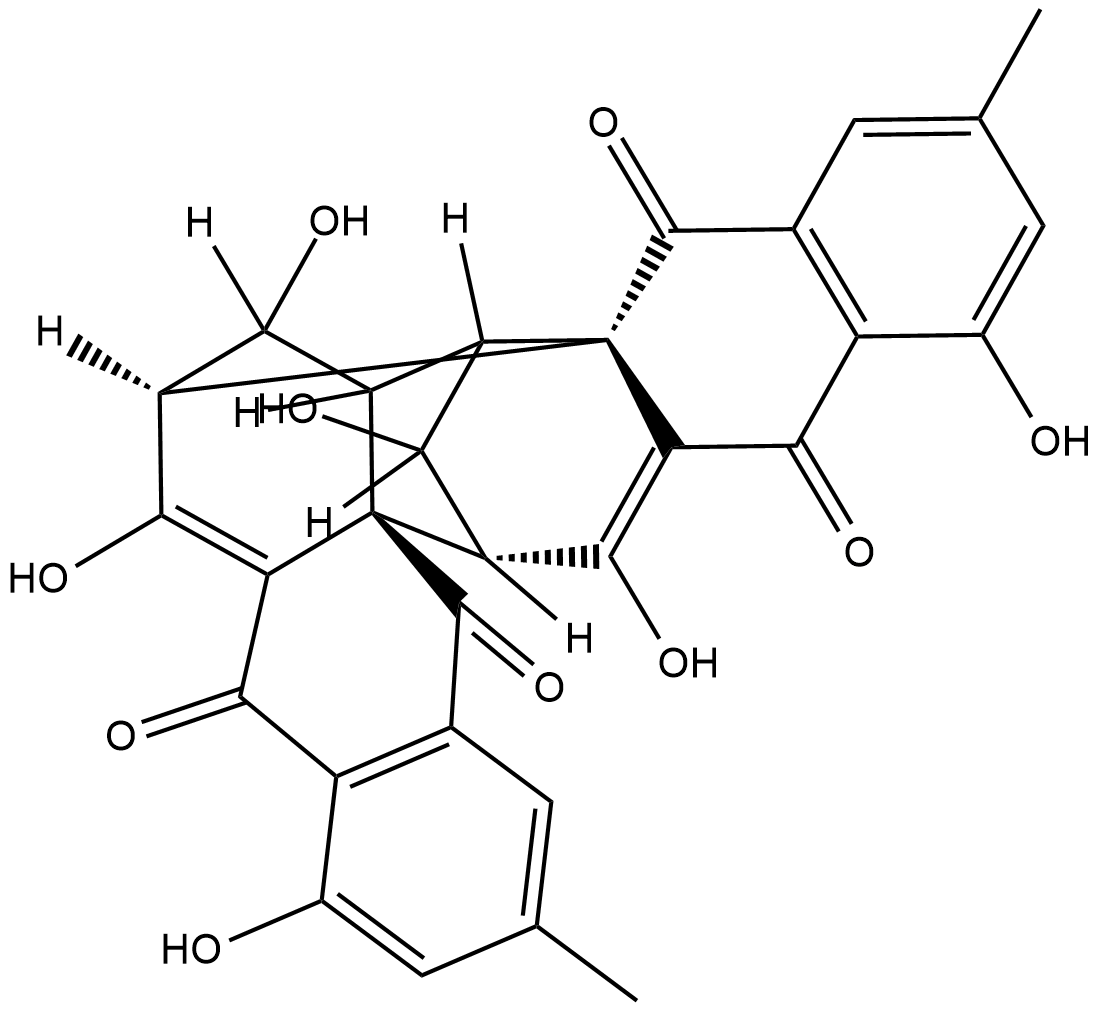
-
GC63969
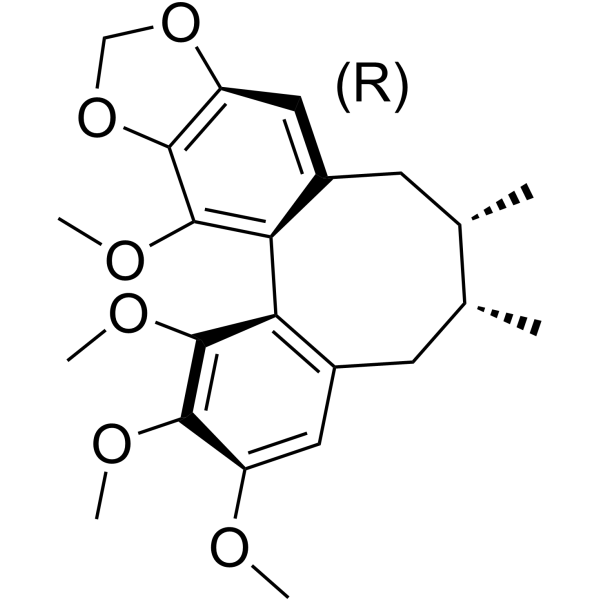
(+)-Schisandrin B
(+)-Schisandrin B ist ein Enantiomer von Schisandrin B.
-
GC40264
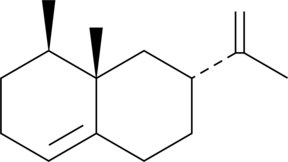
(+)-Valencene
NSC 148969
(+)-Valencene is a sesquiterpene that has been found in C.
-
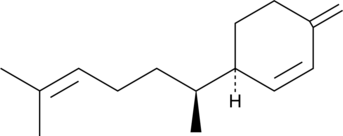
GC49502
(-)-β-Sesquiphellandrene
A sesquiterpene with antiviral and anticancer activities
-
GC32705
(-)-DHMEQ (Dehydroxymethylepoxyquinomicin)
Dehydroxymethylepoxyquinomicin
(-)-DHMEQ (Dehydroxymethylepoxychinomicin) (Dehydroxymethylepoxychinomicin) ist ein potenter, selektiver und irreversibler NF-⋺B-Inhibitor, der sich kovalent an einen Cysteinrest bindet.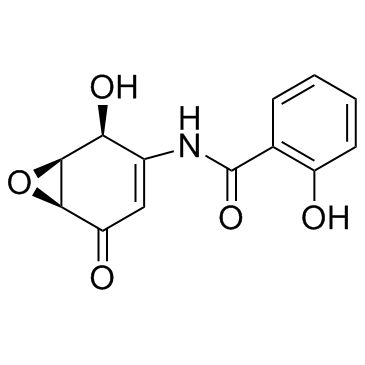
-
GC14049
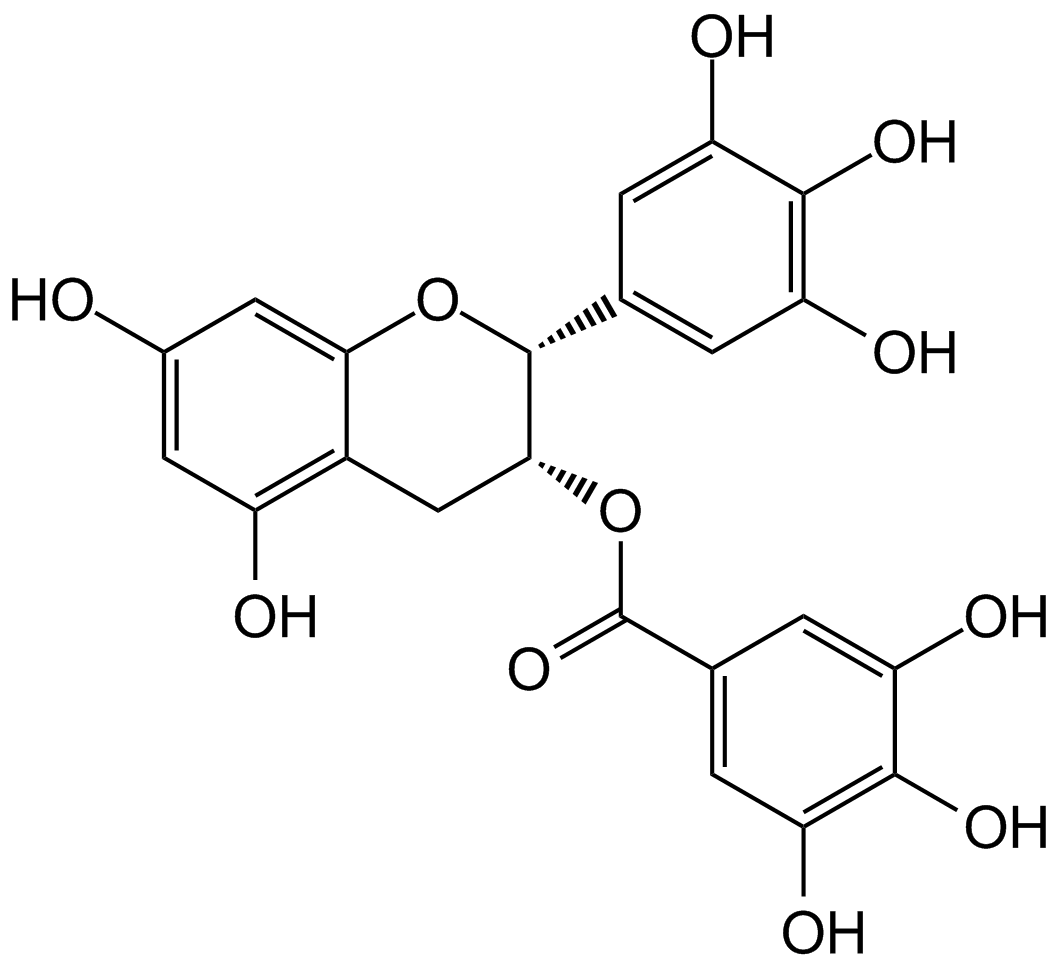
(-)-Epigallocatechin gallate (EGCG)
EGCG
Ein Phenol mit vielfältigen biologischen Aktivitäten.
-
GC45248
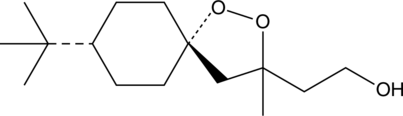
(-)-FINO2
(-)-FINO2 ist ein potenter Ferroptose-Induktor. (-)-FINO2 hemmt die AktivitÄt von GPX4. (-)-FINO2 ist ein stabiles Oxidationsmittel, das Eisen oxidiert und bei unterschiedlichen pH-Werten stabil ist. (-)-FINO2 verursacht eine weit verbreitete Lipidperoxidation.
-
GC46245
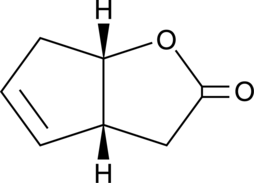
(-)-G-Lactone
A bicyclic γ-lactone
-
GC38316
(-)-Limonene
(±)-Dipentene, DL-Limonene, NSC 844, NSC 21446
(-)-Limonen ((S)-(-)-Limonen) ist ein Monoterpen, das in vielen KiefernnadelÖlen und in Terpentin vorkommt.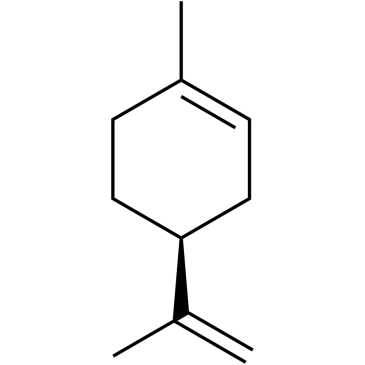
-
GC46247
(-)-Mycousnine
Mycousunin
A microbial metabolite with antibacterial and antifungal activities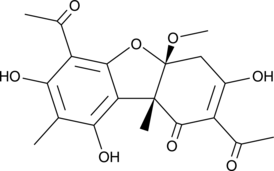
-
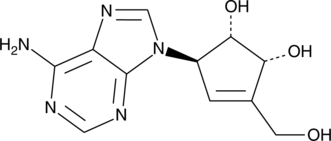
GC45251
(-)-Neplanocin A
S-Adenosylhomocysteine (SAH) hydrolase catalyzes the reversible hydrolysis of SAH to adenosine and homocysteine.
-
GC45272
(-)-Rasfonin
TT-1
(-)-Rasfonin ist ein Pilz-SekundÄrmetabolit und hemmt kleine G-Proteine Ras. (-)-Rasfonin induziert Apoptose, Nekrose und Autophagie in ACHN-Zellen (einer Nierenkarzinom-Zelllinie).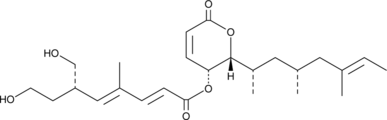
-
GC40803
(25S)-δ7-Dafachronic Acid
UPF1404
During unfavorable environmental conditions, C.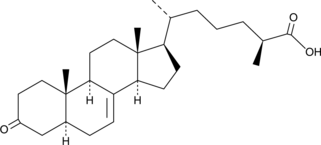
-
GC52442
(D)-PPA 1 (trifluoroacetate salt)
DPPA-1, NYSKPTDRQYHF
An inhibitor of the PD-1-PD-L1 protein-protein interaction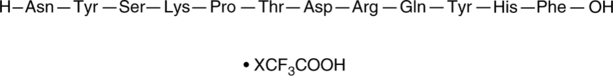
-
GC41700
(E)-2-(2-Chlorostyryl)-3,5,6-trimethylpyrazine
CSTMP
(E)-2-(2-Chlorostyryl)-3,5,6-trimethylpyrazine (CSTMP) is a stilbene derivative with antioxidant and anticancer activities.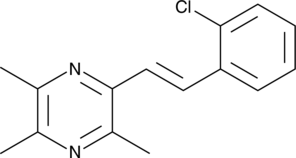
-
GC61668
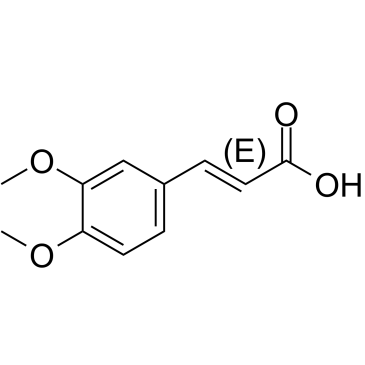
(E)-3,4-Dimethoxycinnamic acid
(E)-3,4-DimethoxyzimtsÄure ist das weniger aktive Isomer von 3,4-DimethoxyzimtsÄure.
-
GC41702
(E)-5-(2-Bromovinyl)uracil
BVU
(E)-5-(2-Bromovinyl)uracil (BVU) is a pyrimidine base and an inactive metabolite of the antiviral agents sorivudine and (E)-5-(2-bromovinyl)-2'-deoxyuridine (BVDU) that may be regenerated to BVDU in vivo.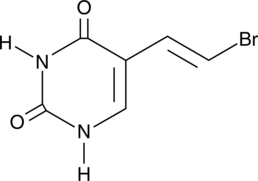
-
GC49003
(E)-Ajoene
NSC 614554
A disulfide with diverse biological activities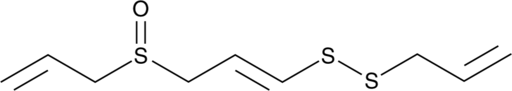
-
GC41703
(E)-C-HDMAPP (ammonium salt)
(E)5hydroxy4methylpent3enyl pyrophosphate
Synthetic and natural alkyl phosphates, also known as phosphoantigens, stimulate the proliferation of γδ-T lymphocytes.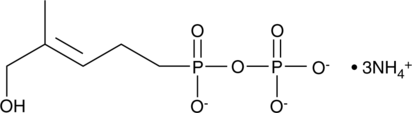
-
GC39747
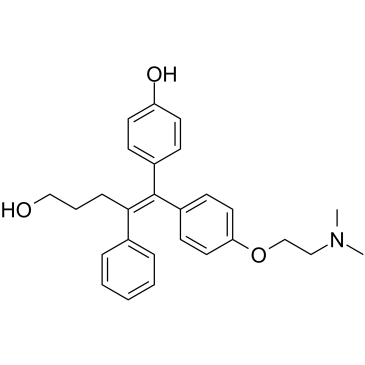
(E/Z)-GSK5182
(E/Z)-GSK5182 ist eine racemische Verbindung der (E)-GSK5182- und (Z)-GSK5182-Isomere.
-
GC61564
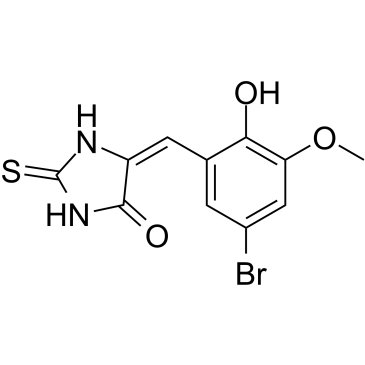
(E/Z)-IT-603
(E/Z)-IT-603 ist eine Mischung aus E-IT-603 und Z-IT-603 (IT-603).
-
GC41721
(R)-α-Lipoic Acid
(R)-(+)-Lipoic Acid
(R)-α-Lipoic acid is the naturally occurring enantiomer of lipoic acid, a cyclic disulfide antioxidant.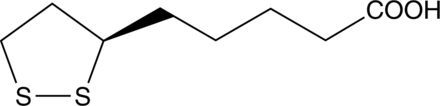
-
GC49167
(R)-(+)-Trityl glycidyl ether
(R)-Trityl Glycidol
A synthetic precursor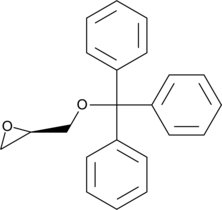
-
GC13030
(R)-(-)-Ibuprofen
(-)-Ibuprofen
(R)-(-)-Ibuprofen ist das R-Enantiomer von Ibuprofen, inaktiv bei COX, hemmt die NF-κB-Aktivierung; (R)-(-)-Ibuprofen zeigt entzÜndungshemmende und antinozizeptive Wirkungen.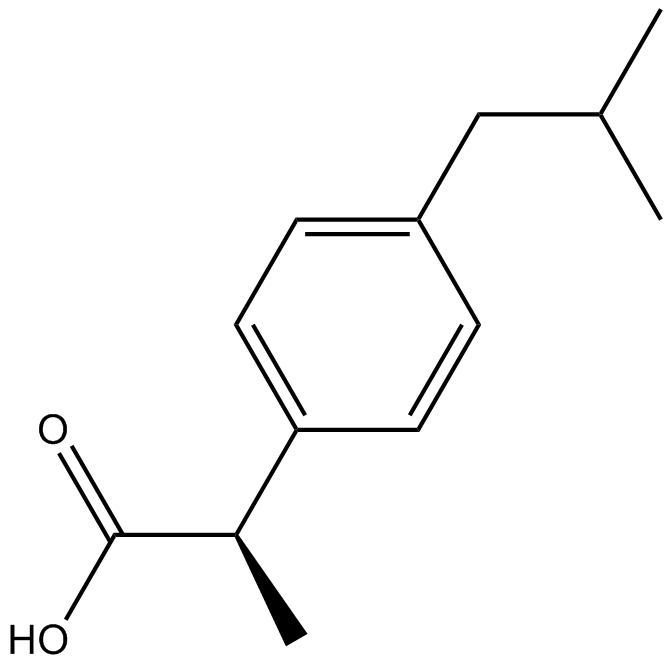
-
GC69823
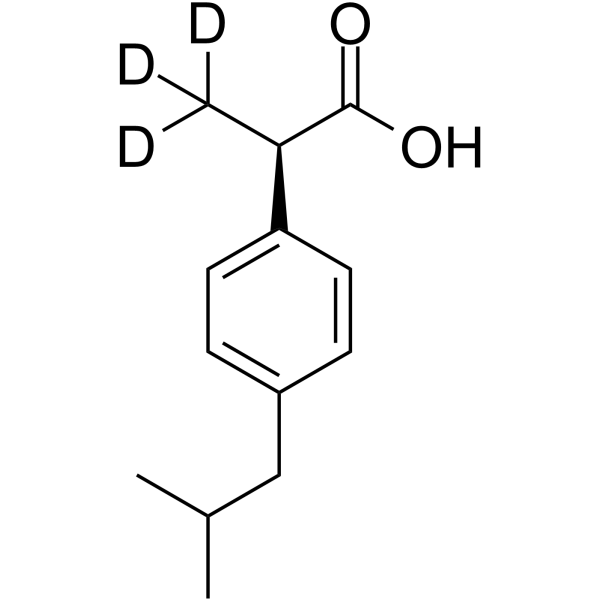
(R)-(-)-Ibuprofen-d3
(R)-Ibuprofen-d3
(R)-(-)-Ibuprofen-d3 ist das Deuterium-Isotop von (R)-(-)-Ibuprofen. (R)-(-)-Ibuprofen ist das R-Isomer von Ibuprofen, hat keine Wirkung auf COX und kann die Aktivierung von NF-κB hemmen. (R)-(-)-Ibuprofen hat entzündungshemmende Eigenschaften und kann zur Erforschung der Schmerzlinderung eingesetzt werden.
-
GC41620
(R)-(-)-Mellein
Ochracin
(R)-(-)-Mellein ist ein aus KulturflÜssigkeiten dieses Aspergillus isoliertes Antibiotikum.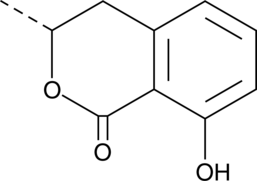
-
GC41712
(R)-3-hydroxy Myristic Acid
(R)-3-hydroxy Tetradecanoic Acid
Lipopolysaccharides (LPS) are components of the cell walls of Gram-negative bacteria.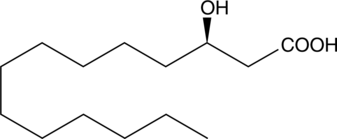
-
GC65610
(R)-5-Hydroxy-1,7-diphenyl-3-heptanone
(R)-5-Hydroxy-1,7-diphenyl-3-heptanon ist ein Diarylheptanoid, das in Alpinia officinarum vorkommt.

-
GC65373
(R)-IL-17 modulator 4
(R)-IL-17-Modulator 4 ist die R-Konfiguration von IL-17-Modulator 4.

-
GC12578
(R)-Lisofylline
(−)-Lisofylline,(R)-LSF
(R)-Lisofyllin ((R)-Lisophyllin) ist ein (R)-Enantiomer des Metaboliten von Pentoxifyllin mit entzÜndungshemmenden Eigenschaften.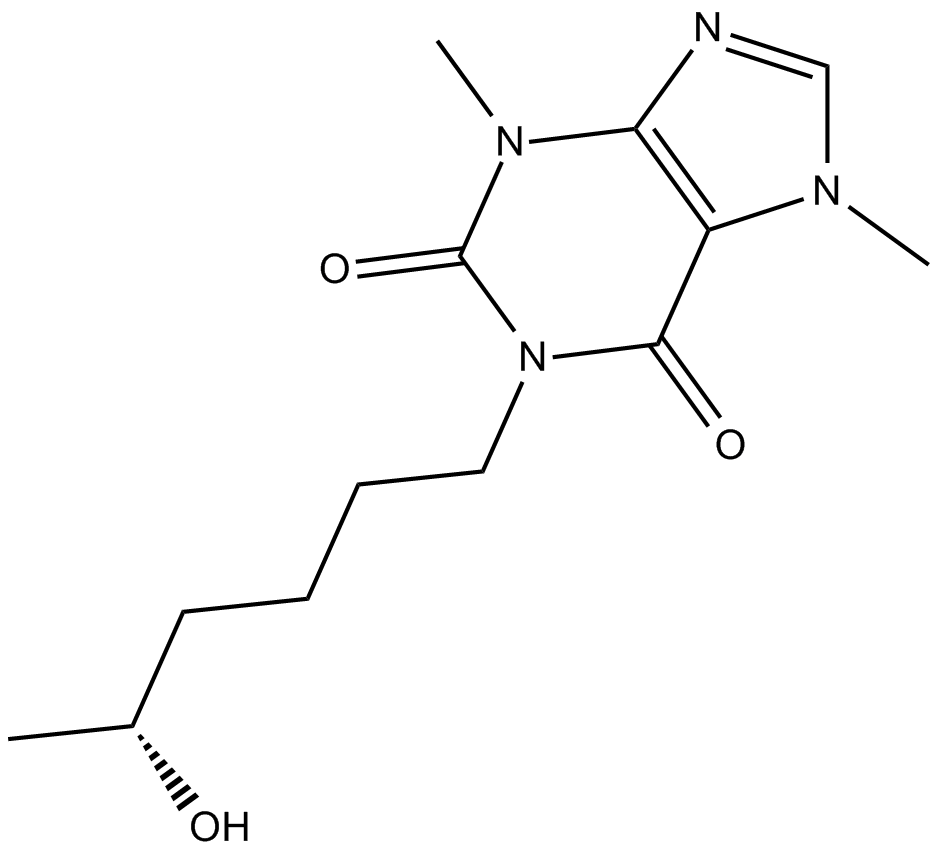
-
GC52185
(R,S)-Anatabine-d4
(±)-Anatabine-d4

-
GC39321
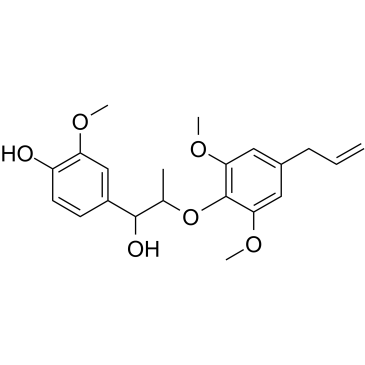
(Rac)-Myrislignan
(Rac)-Myrislignan ist das Racemat von Myrislignan.
-
GC66334
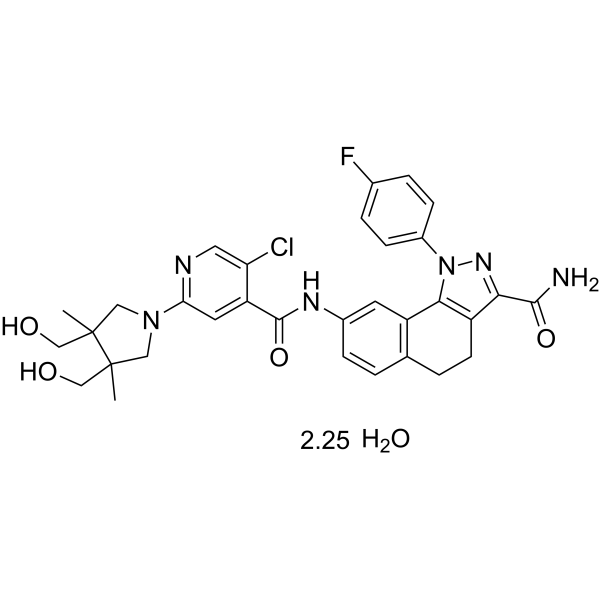
(Rac)-PF-184 hydrate
(Rac)-PF-184-Hydrat ist ein potenter Inhibitorfaktor-⋺B-Kinase-2 (IKK-2)-Inhibitor mit einem IC50 von 37 nM. (Rac)-PF-184-Hydrat wirkt entzÜndungshemmend.
-
GC69799
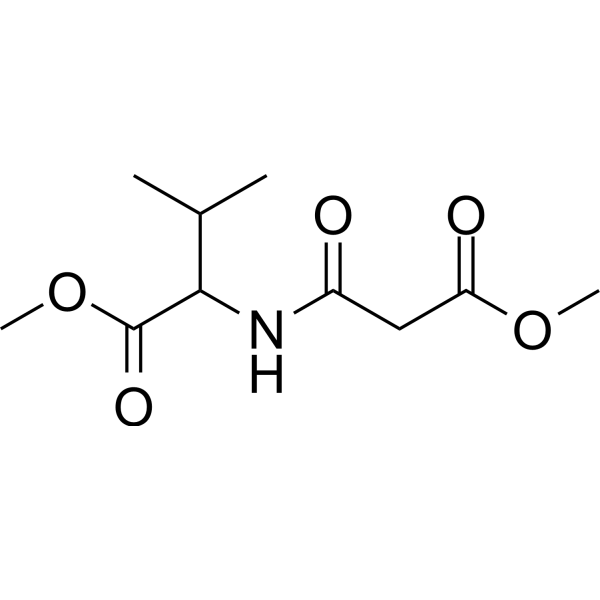
(Rac)-ZLc-002
(Rac)-ZLc-002 ist ein Inhibitor, der die Interaktion zwischen nNOS und dem Protein NOS1AP hemmt. Er hemmt entzündungsbedingte Schmerzen und neuropathische Schmerzen, die durch Chemotherapie verursacht werden, und senkt in Kombination mit Paclitaxel die Aktivität von Tumorzellen.
-
GC46345
(S)-(-)-Perillaldehyde
(–)-Perillaldehyde, L-Perillaldehyde, (S)-Perillaldehyde
(S)-(-)-Perillaldehyd ist ein Hauptbestandteil des in Perillae Herba enthaltenen Ätherischen Öls.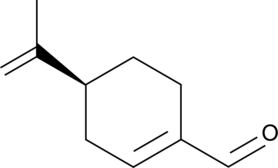
-
GC49028
(S)-3-Thienylglycine
L-R-(3-Thienyl)glycine, L-α-3-Thienylglycine
A thienyl-containing amino acid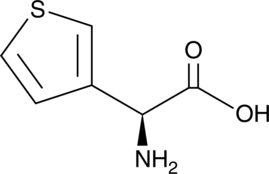
-
GC52192
(S)-4'-nitro-Blebbistatin
(-)-4'-nitro-Blebbistatin, p-nitro-Blebbistatin, para-nitro-Blebbistatin
(S)-4'-Nitro-Blebbistatin ist ein nicht zytotoxischer, photostabiler, fluoreszierender und spezifischer Myosin-II-Inhibitor, der in der Untersuchung der spezifischen Rolle von Myosin II in physiologischen, entwicklungsbezogenen und zellbiologischen Studien verwendet wurde.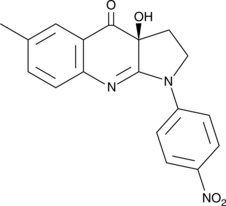
-
GC48719
(S)-Canadine
(–)-Canadine, (S)-Tetrahydroberberine
(S)-Canadine ist ein Alkaloid und Zwischenprodukt in der Biosynthese von Berberin mit insektizider Wirkung.
-
GC46352
(S)-DO271
An inactive control for DO264